DOI: https://doi.org/10.1038/s41368-024-00294-z
PMID: https://pubmed.ncbi.nlm.nih.gov/38719817
تاريخ النشر: 2024-05-08
التقسيم التلقائي الكامل بالذكاء الاصطناعي للأنسجة المتعلقة بجراحة الفم استنادًا إلى صور الأشعة المقطعية المخروطية
الملخص
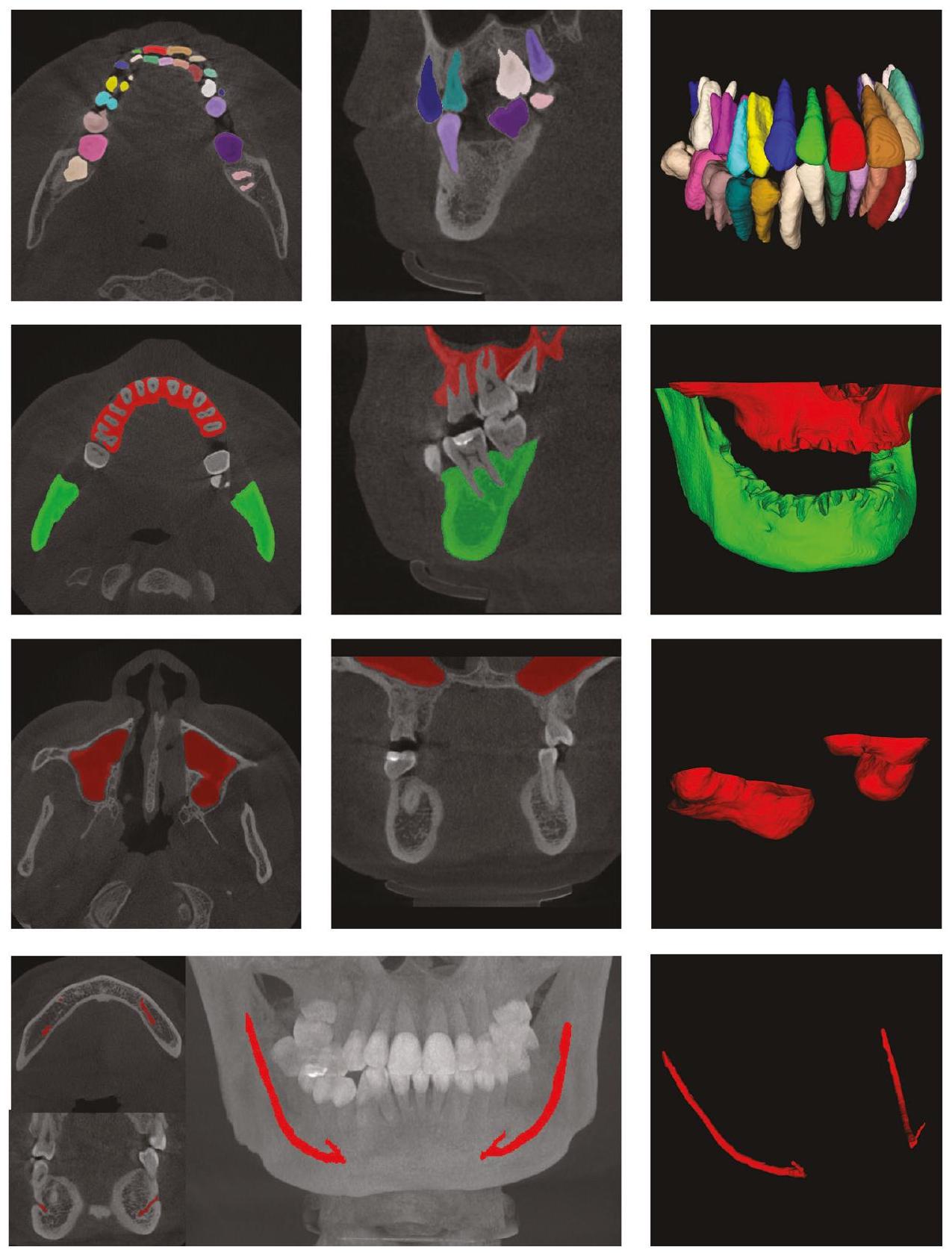
يمكن أن يسرع التقسيم الدقيق للأنسجة المتعلقة بجراحة الفم من صور الأشعة المقطعية المخروطية (CBCT) بشكل كبير من تخطيط العلاج ويحسن دقة الجراحة. في هذه الورقة، نقترح نظام تقسيم أنسجة مؤتمت بالكامل لجراحة زراعة الأسنان. على وجه التحديد، نقترح طريقة معالجة مسبقة للصور تعتمد على توزيعات البيانات، والتي يمكن أن تعالج صور CBCT بشكل تكيفي مع معلمات مختلفة. بناءً على ذلك، نستخدم شبكة تقسيم العظام للحصول على نتائج تقسيم العظم السنخي، والأسنان، والجيوب الأنفية العلوية. نستخدم مناطق الأسنان والفك السفلي كمناطق ROI لتقسيم الأسنان وتقسيم أنبوب العصب الفك السفلي لتحقيق المهام المقابلة. يمكن أن توفر نتائج تقسيم الأسنان معلومات الترتيب للأسنان. تظهر النتائج التجريبية المقابلة أن طريقتنا يمكن أن تحقق دقة وكفاءة تقسيم أعلى مقارنة بالطرق الحالية. كانت متوسط درجات Dice لدينا في مهام تقسيم الأسنان، والعظم السنخي، والجيوب الأنفية العلوية، وقناة الفك السفلي
مقدمة
تقييم دقيق بعد الجراحة،
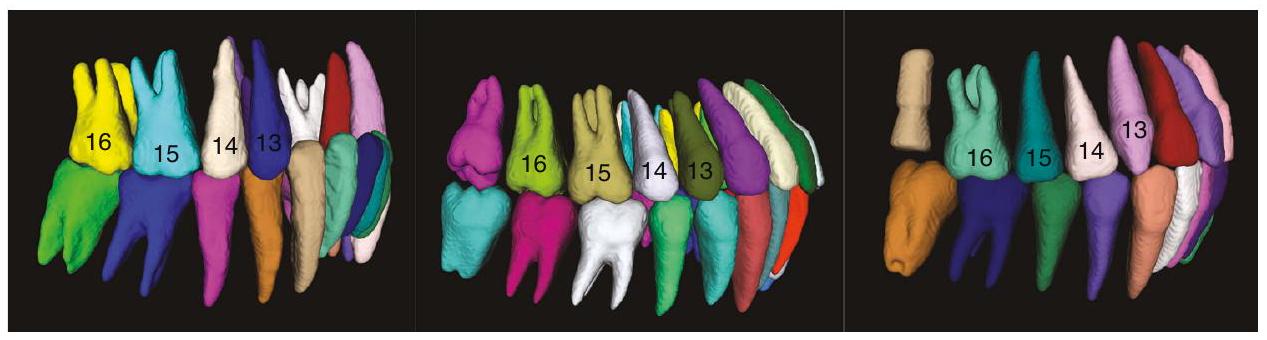
- نتائج تقسيم الأسنان للطرق الحالية لا تحدد الأسنان وفقًا لتدوين FDI ذو الرقمين.
حتى لا يمكن اكتشاف المعلومات المفقودة عن الأسنان بدقة وتمثيلها. لقد قمنا بإعادة إنتاج طريقة كوي وآخرون. الذي يُعتبر الأحدث في هذا المجال، تظهر النتيجة في الشكل 1. لا يمكنه تحقيق نفس نتائج التصنيف للأسنان في نفس الموضع، والذي تم تمثيله في الشكل بألوان مختلفة للأسنان في نفس الموضع. الطريقة التي اقترحها ليو وآخرون يتطلب ذلك أولاً تقسيم نموذج المسح الفموي وتسجيله مع صورة CBCT للحصول على نتائج التقسيم. هذا يحد من تطبيق هذه الطرق في تحليل وتشخيص العيوب السنية، حيث أن المعلومات حول الأسنان المفقودة ضرورية للتحديد الصحيح وتقييم حالة صحة الفم لدى المريض. - على عكس الأشعة المقطعية التقليدية، تستخدم الشركات المصنعة لمعدات الأشعة المقطعية المخروطية (CBCT) أجهزة تصوير وأساليب استيفاء مختلفة، كما تختار ديناميكيًا معلمات التصوير وفقًا لحالة المريض أثناء الاستخدام، مما يؤدي إلى أن صور CBCT ستحتوي على توزيع تدرج رمادي وتباين مختلف، مما يجعل من الصعب تطبيق صورة بسيطة.
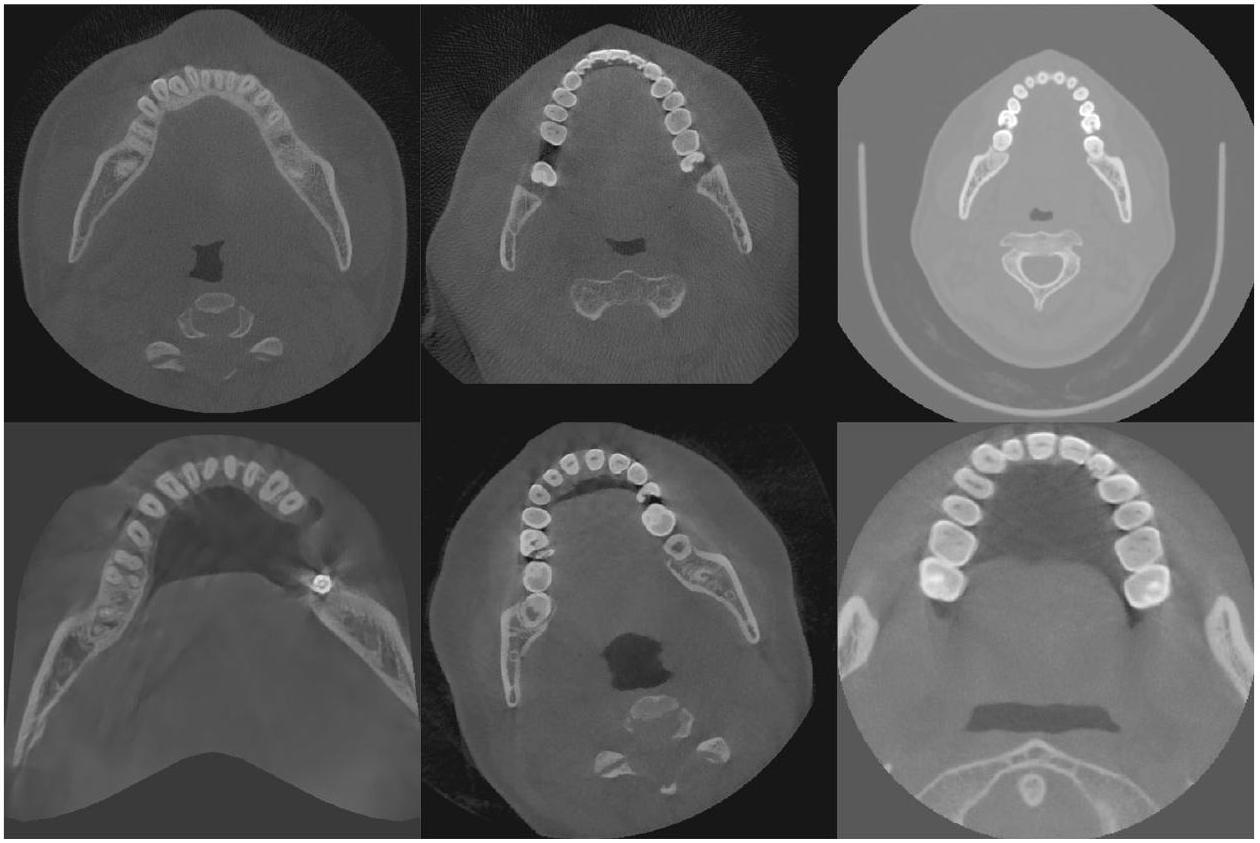
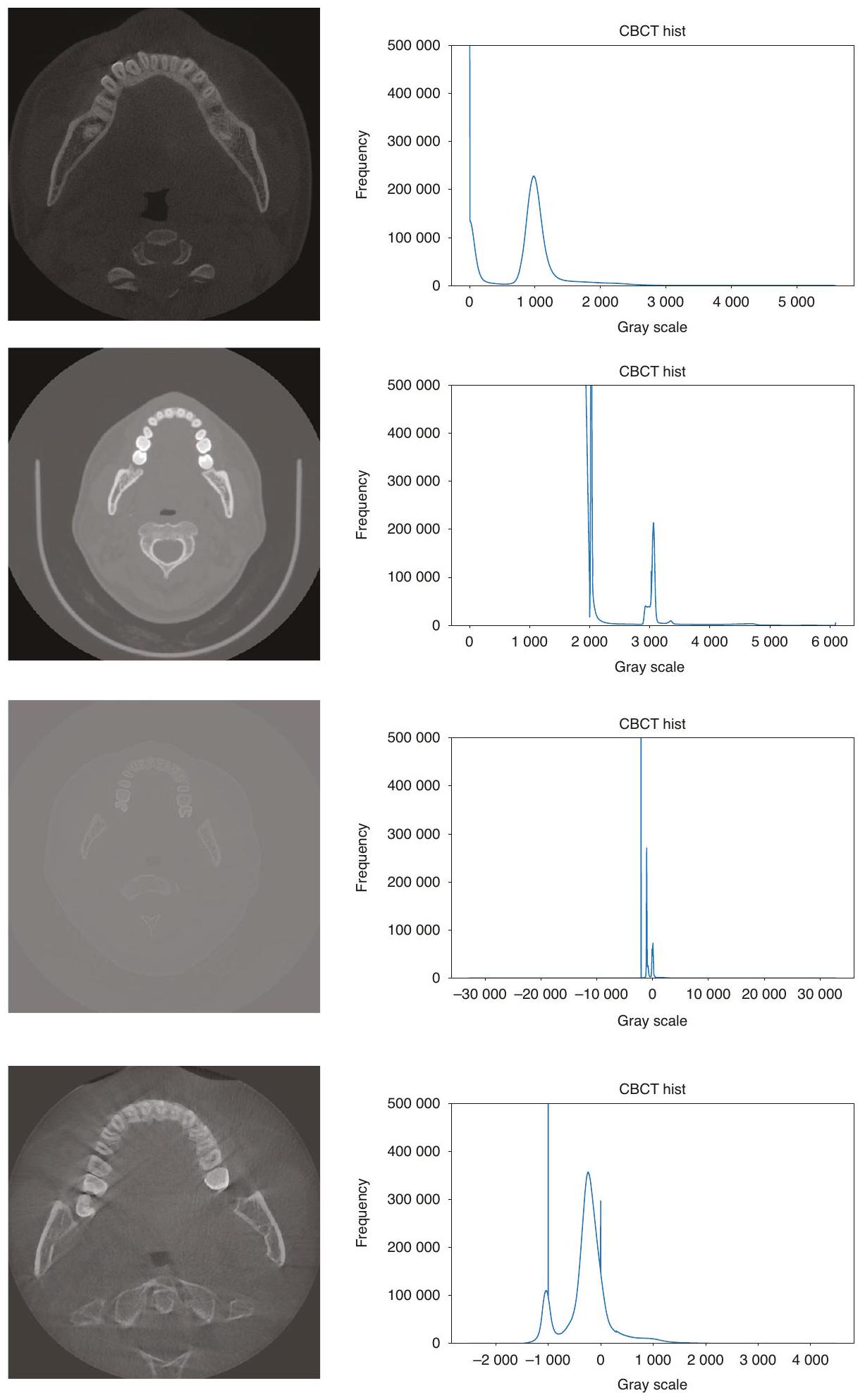
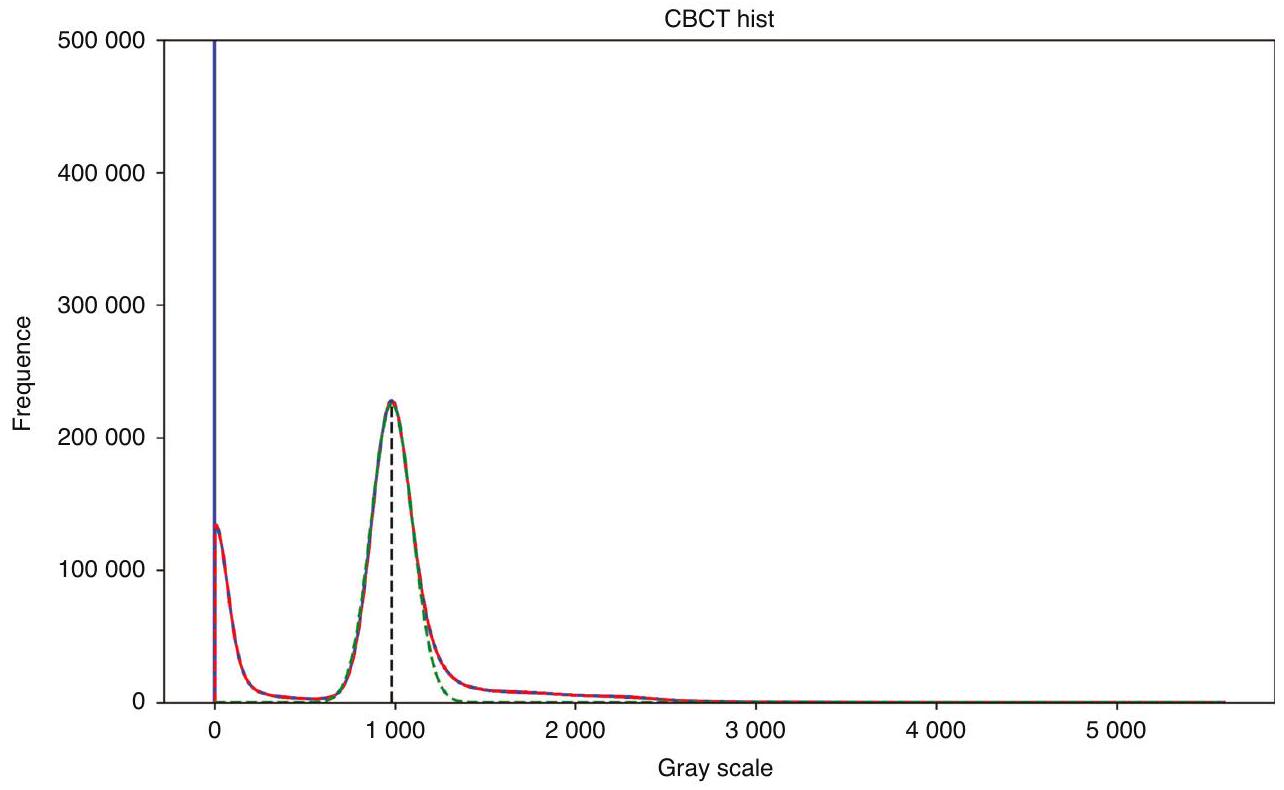
طرق المعالجة المسبقة لجميع صور CBCT؛ وفي مهمة تقسيم الأسنان، فإن معظم الصور الأصلية هي مناطق أنسجة لينة غير مفيدة للتقسيم، مما يؤدي إلى توزيع غير متوازن للبيانات، وهو ما لا تعالجه الطرق الحالية. تُظهر الصور الفعلية لـ CBCT في الشكل 2، ويمكن ملاحظة وجود اختلافات كبيرة في تباين الصورة، ومجال الرؤية، وما إلى ذلك. - الطرق الحالية معقدة للغاية وغالبًا ما تتطلب خطوات متعددة للحصول على نتائج تقسيم جيدة. قد تتطلب هذه الطرق استخدام تقنيات معالجة مسبقة متعددة، وطرق استخراج الميزات، والمصنفات أو النماذج، وخطوات المعالجة اللاحقة لإكمال تقسيم الأسنان. تؤدي هذه التعقيدات إلى طرق أقل كفاءة من الناحية الحسابية، حيث قد يقدم كل خطوة أخطاء أو عيوب، وكل خطوة في العملية الكاملة تتطلب ضبطًا دقيقًا وتحققًا، مما يزيد من صعوبة تطوير الطرق وتطبيقها.
- الطرق الحالية لا تؤدي جميع مهام التقسيم هذه بشكل تلقائي بالكامل بطريقة شاملة، حيث تركز عادةً على مهمة واحدة، مثل تقسيم الأسنان أو تقسيم العظم السنخي في منطقة محددة مسبقًا (ROI)، مع القليل من البحث حول تقسيم القناة الفكية من الجيب الفكي.
النتائج

| فصول القطاعات | نرد/% | ملاو/% | HD/mm | ASD/mm |
| سن |
|
|
|
|
| عظم الفك العلوي |
|
|
|
|
| عظم الفك |
|
|
|
|
| القناة الفكية |
|
|
|
|
| الجيوب الأنفية الفكية |
|
|
|
|
| متوسط |
|
|
|
|
| الجدول 2. النتائج التجريبية للمقارنة الكمية مع الأساليب المتقدمة الحالية من حيث دقة التقسيم والكشف | ||||
| طرق | نرد/% | ملاو/% | HD/mm | ASD/mm |
| هاي-مو توث سيج |
|
|
|
|
| nnUNet |
|
|
|
|
| توث نت |
|
|
|
|
| ريلو-نت |
|
|
|
|
| دينس أيه إس بي بي – يونيك |
|
|
|
|
| خاصتنا |
|
|
|
|
| النص العريض يمثل أعلى قيمة في عموده | ||||
| الجدول 3. تحليل تأثير اختيار علاقات مختلفة بين
|
||||
| عناصر | نرد/% | ملاو/% | HD/mm | ASD/mm |
| معالجة مسبقة عادية |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| النص العريض يمثل أعلى قيمة في عموده | ||||
نقاش
| الطرق | T1 | T2 | T3 | T4 | T5 | T6 | T7 | T8 |
| طريقتنا |
|
|
|
|
|
|
|
|
| الشركة المصنعة | اسم طراز الشركة المصنعة | Dice/% | mloU/% | HD/mm | ASD/mm |
| Carestream Health | CS 9300، CS 9301 |
|
|
|
|
| Imaging Sciences International | 9-17 |
|
|
|
|
| J.Morita.Mfg.Corp. |
|
|
|
|
|
| LargeV | HighRes3D، SMART3D |
|
|
|
|
| NewTom | NTVGiMK4، NTVGiEVO، NT5G |
|
|
|
|
| NNT | NTVGiEVO |
|
|
|
|
| PaloDEx Group Oy | ORTHOPANTOMOGRAPH OP 3D |
|
|
|
|
| RAY Co., Ltd. | RAYSCAN N Alpha Plus |
|
|
|
|
| Sirona | ORTHOPHOS SL |
|
|
|
|
| Vatech Company Limited | PHT-35LHS |
|
|
|
|
| YOFO | Pirox-R |
|
|
|
|
| العناصر | الذكاء الاصطناعي | الخبير | الذكاء الاصطناعي المساعد + الضبط اليدوي |
| الوقت
|
|
240 | 5.6 |
| Dice/% |
|
92.6 | |
| النص الغامق يمثل أعلى قيمة في عموده | |||
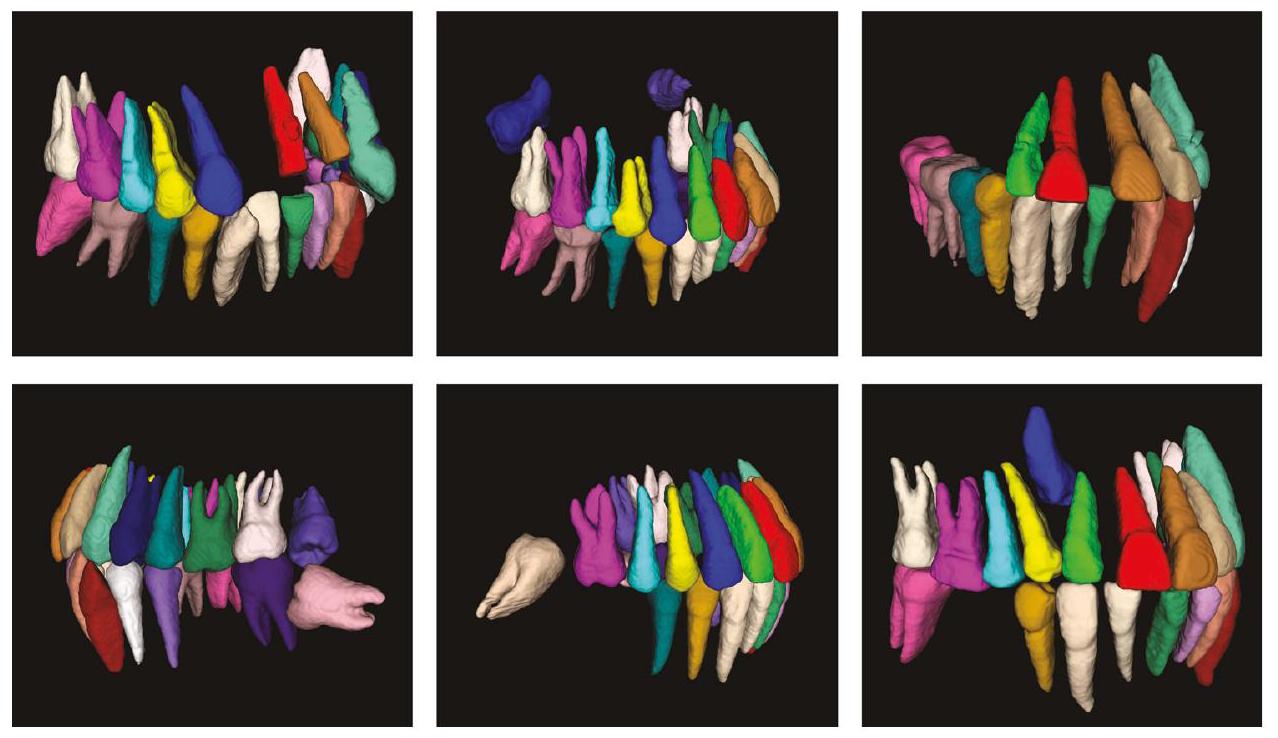
الخوارزمية صعبًا ويجعل من المستحيل تحديد واختيار أسنان معينة بدقة أثناء التفاعل مع البرمجيات. إن القدرة على التقاط المعلومات بدقة حول الأسنان المفقودة تجعل طريقتنا أكثر ملاءمة للتقييم الشامل للأسنان.
استنتاج
طرق البحث
طريقة المعالجة التكيفية لصورة CBCT
هيكل الشبكة
من الصعب الحصول على العلاقة بين سنين بعيدتين عند إجراء تجزئة الأسنان، مما يؤدي إلى انخفاض دقة التجزئة. تتطلب طرق تجزئة الأسنان المعتمدة على الشبكات العصبية التلافيفية الحالية معلومات ميزات أخرى كمدخلات، مثل مراكز الأسنان التي تم الحصول عليها مسبقًا، ومواقع الأسنان التي تم الحصول عليها باستخدام نماذج المسح الفموية، إلخ.

نموذج العلاقات بين البكسلات البعيدة واستخراج المعلومات المحلية في نفس الوقت، وهو أمر حاسم لتوقع موضع الأسنان.
Swin-Transformer: كمستخرج ميزات، يستخدم لاستخراج تمثيلات ميزات ذات مغزى من الصورة المدخلة. يعتمد على بنية Swin-Transformer
| أبعاد التضمين | حجم الميزة | عدد الكتل | حجم النافذة | عدد الرؤوس | المعلمات | FLOPS |
| 768 | 48 |
|
|
|
62.19 M | 394.84 G |
| الشركة المصنعة | اسم طراز الشركة المصنعة | الجنس (أنثى/ ذكر) | جهد الأنبوب/kVp | تيار الأنبوب
|
المسافة/ مم | متوسط العمر/ سنوات | عدد CBCT (الحالات) |
| Carestream Health | CS 9300، CS 9301 | 90 | 10 | 0.18 | 40.1 | 15 | |
| دولي | |||||||
| J.Morita.Mfg.Corp. | 2F/2M | 89 | 7 | 0.25 | 25.3 | 4 | |
| LargeV | HighRes3D، SMART3D |
|
100 | 4 | 0.25 | 44.5 | 112 |
| NewTom | NTVGiMK4، NTVGiEVO، NT5G | 22F/13M | 110 | 1، 2، 3، 4، 5، 7، 9، 10 | 0.3، 0.25 | 42.1 | 35 |
| NNT | NTVGiEVO | 7M | 110 | 3، 7، 8، 9، 10، 11، 14 | 0.3 | 31.2 | 7 |
| PaloDEx Group Oy | ORTHOPANTOMOGRAPH OP 3D | 5M | 95 | 3، 8 | 0.25 | 44.2 | 7 |
| RAY Co.، Ltd. | RAYSCAN N Alpha Plus | 1F/2M | 90 | 10 | 1 | 3 | |
| Sirona | ORTHOPHOS SL | 85 | 10 | 0.22 | 46.7 | 10 | |
| Vatech Company Limited | PHT-35LHS | 1F/3M | 94 | 8 | 0.2 | 47.8 | 4 |
| YOFO | Pirox-R | 29F/23M | 90 | 8 | 0.25 | 40.1 | 52 |
| 0.4 | 91 | ||||||
| 0.4 | 97 | ||||||
دالة الخسارة
على التوالي، والتي تم تعيينها إلى 1 في هذه التجربة.
مجموعة البيانات
معايير التقييم
تفاصيل التنفيذ
شكر وتقدير
مساهمات المؤلفين
معلومات إضافية
REFERENCES
- Bai, S. Z. et al. [Animal experiment on the accuracy of the Autonomous Dental Implant Robotic System]. Zhonghua Kou Qiang Yi Xue Za Zhi 56, 170-174 (2021).
- Jia, S., Wang, G., Zhao, Y. & Wang, X. Accuracy of an autonomous dental implant robotic system versus static guide-assisted implant surgery: a retrospective clinical study. J. Prosthet. Dent. https://doi.org/10.1016/j.prosdent.2023.04.027 (2023).
- Li, Z., Xie, R., Bai, S. & Zhao, Y. Implant placement with an autonomous dental implant robot: a clinical report. J. Prosthet. Dent. https://doi.org/10.1016/ j.prosdent.2023.02.014 (2023).
- Wu, Q., Zhao, Y. M., Bai, S. Z. & Li, X. Application of robotics in stomatology. Int. J. Comput. Dent. 22, 251-260 (2019).
- Cheng, K. J. et al. Accuracy of dental implant surgery with robotic position feedback and registration algorithm: an in-vitro study. Comput. Biol. Med. 129, 104153 (2021).
- Tao, B. et al. Accuracy of dental implant surgery using dynamic navigation and robotic systems: an in vitro study. J. Dent. 123, 104170 (2022).
- Bolding, S. L. & Reebye, U. N. Accuracy of haptic robotic guidance of dental implant surgery for completely edentulous arches. J. Prosthet. Dent. 128, 639-647 (2022).
- Yang, X. et al. ImplantFormer: vision transformer based implant position regression using dental CBCT data. Accessed July 3, 2023. Preprint at http:// arxiv.org/abs/2210.16467 (2023).
- Płotka, S. et al. Convolutional neural networks in orthodontics: a review. Accessed July 3, 2023. Preprint at http://arxiv.org/abs/2104.08886 (2021).
- Duy, N. T., Lamecker, H., Kainmueller, D. & Zachow, S. Automatic detection and classification of teeth in CT data. In Medical Image Computing and ComputerAssisted Intervention – MICCAI 2012, Lecture Notes in Computer Science (eds Ayache, N., Delingette, H., Golland, P. & Mori, K.) 609-616 (Springer, 2012).
- Barone, S., Paoli, A. & Razionale, A. V. CT segmentation of dental shapes by anatomy-driven reformation imaging and B-spline modelling. Int. J. Numer. Method. Biomed. Eng. 32, e02747 (2016).
- Sepehrian, M., Deylami, A. M. & Zoroofi, R. A. Individual teeth segmentation in CBCT and MSCT dental images using watershed. In 2013 20th Iranian Conference on Biomedical Engineering (ICBME) 27-30 (IEEE, 2013).
- Gao, H. & Chae, O. Individual tooth segmentation from CT images using level set method with shape and intensity prior. Pattern Recognit. 43, 2406-2417 (2010).
- Qian, J. et al. An automatic tooth reconstruction method based on multimodal data. J. Vis. 24, 205-221 (2021).
- Poonsri, A., Aimjirakul, N., Charoenpong, T. & Sukjamsri, C. Teeth segmentation from dental X-ray image by template matching. In 2016 9th Biomedical Engineering International Conference (BMEICON) 1-4 (IEEE, 2016).
- Akhoondali, H., Zoroofi, R. A. & Shirani, G. Rapid automatic segmentation and visualization of teeth in CT-scan data. J. Appl. Sci. 9, 2031-2044 (2009).
- Zou, X., Liu, W., Wang, J., Tao, R. & Zheng, G. ARST: auto-regressive surgical transformer for phase recognition from laparoscopic videos. Accessed July 3, 2023. Preprint at http://arxiv.org/abs/2209.01148 (2022).
- Huang, S., Xu, T., Shen, N., Mu, F. & Li, J. Rethinking few-shot medical segmentation: a vector quantization view.
- Zhang, J., Xie, Y., Xia, Y. & Shen, C. DoDNet: learning to segment multi-organ and tumors from multiple partially labeled datasets. Accessed July 3, 2023. Preprint at http://arxiv.org/abs/2011.10217 (2020).
- Wang, R. et al. Medical image segmentation using deep learning: a survey. IET Image Process. 16, 1243-1267 (2022).
- Zhang, R. et al. Multiple supervised residual network for osteosarcoma segmentation in CT images. Comput. Med. Imaging Graph. 63, 1-8 (2018).
- Tao, R., Liu, W. & Zheng, G. Spine-transformers: vertebra labeling and segmentation in arbitrary field-of-view spine CTs via 3D transformers. Med. Image Anal. 75, 102258 (2022).
- Cai, Y. et al. Swin Unet3D: a three-dimensional medical image segmentation network combining vision transformer and convolution. BMC Med. Inf. Decis. Mak. 23, 33 (2023).
- Cao, H. et al. Swin-Unet: Unet-Like Pure Transformer for Medical Image Segmentation. In Computer Vision – ECCV 2022 Workshops. ECCV 2022. Lecture Notes in Computer Science, Vol. 13803 (eds. Karlinsky, L., Michaeli, T. & Nishino, K.) https://doi.org/10.1007/978-3-031-25066-8_9 (Springer, Cham, 2023).
12
25. Hatamizadeh, A. et al. Swin UNETR: Swin Transformers for Semantic Segmentation of Brain Tumors in MRI Images. https://arxiv.org/abs/2201.01266 (2022).
26. Milletari, F., Navab, N. & Ahmadi, S.-A. V-Net: Fully Convolutional Neural Networks for Volumetric Medical Image Segmentation. 2016 Fourth International Conference on 3D Vision (3DV), Stanford, CA, USA. 565-571, https://doi.org/10.1109/3DV.2016.79 (2016).
27. Cui, Z. et al. A fully automatic AI system for tooth and alveolar bone segmentation from cone-beam CT images. Nat. Commun. 13, 2096 (2022).
28. Chen, Y. et al. Automatic segmentation of individual tooth in dental CBCT images from tooth surface map by a multi-task FCN. IEEE Access 8, 97296-97309 (2020).
29. Jaskari, J. et al. Deep learning method for mandibular canal segmentation in dental cone beam computed tomography volumes. Sci. Rep. 10, 5842 (2020).
30. Verhelst, P. J. et al. Layered deep learning for automatic mandibular segmentation in cone-beam computed tomography. J. Dent. 114, 103786 (2021).
31. Chung, M. et al. Pose-aware instance segmentation framework from cone beam CT images for tooth segmentation. Comput. Biol. Med. 120, 103720 (2020).
32. Cui, Z., Li, C. & Wang, W. ToothNet: automatic tooth instance segmentation and identification from cone beam CT Images. In 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 6361-6370 (IEEE, 2019).
33. Lee, J. et al. Tooth instance segmentation from cone-beam CT images through point-based detection and Gaussian disentanglement. Multimed Tools Appl. 81, 18327-18342 (2022).
34. Gerhardt, M. D. N. et al. Automated detection and labelling of teeth and small edentulous regions on cone-beam computed tomography using convolutional neural networks. J. Dent. 122, 104139 (2022).
35. Liu, J. et al. Deep learning-enabled 3D multimodal fusion of cone-beam CT and intraoral mesh scans for clinically applicable tooth-bone reconstruction. Patterns 4, 100825 (2023).
36. Nackaerts, O. et al. Segmentation of trabecular jaw bone on cone beam CT datasets: segmentation of jaw bone on CBCT datasets. Clin. Implant Dent. Relat. Res. 17, 1082-1091 (2015).
37. FDI World Dental Federation. Accessed July 11, 2023. https://web.archive.org/web/ 20070401074213/http://www.fdiworldental.org/resources/5_Onotation.html (2007).
38. Cui, Z., et al. Hierarchical morphology-guided tooth instance segmentation from CBCT images. In Information Processing in Medical Imaging, Lecture Notes in Computer Science (eds Feragen, A., Sommer, S., Schnabel, J. & Nielsen, M.) 150-162 (Springer International Publishing, 2021).
39. Isensee, F., Jäger, P. F., Kohl, S. A. A., Petersen, J. & Maier-Hein, K. H. Automated design of deep learning methods for biomedical image segmentation. Nat. Methods 18, 203-211 (2021).
40. Wu, X. et al. Center-sensitive and boundary-aware tooth instance segmentation and classification from cone-beam CT. In 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI) 939-942 (IEEE, 2020).
41. Zhang, Y. & Yu, H. Convolutional neural network based metal artifact reduction in X-ray computed tomography. IEEE Trans. Med. Imaging 37, 1370-1381 (2018).
42. Liang, X. et al. Generating synthesized computed tomography (CT) from conebeam computed tomography (CBCT) using CycleGAN for adaptive radiation therapy. Phys. Med. Biol. 64, 125002 (2019).
43. Liu, Y. et al. CBCT-based synthetic CT generation using deep-attention cycleGAN for pancreatic adaptive radiotherapy. Med. Phys. 47, 2472-2483 (2020).
44. Galibourg, A. et al. Assessment of automatic segmentation of teeth using a watershed-based method. Dentomaxillofac. Radiol. 47, 20170220 (2018).
45. Hounsfield, G. N. Computed medical imaging. Science 210, 22-28 (1980).
46. Özgün, Ç., Abdulkadir, A., Lienkamp, S. S., Brox, T. & Ronneberger, O. 3D U-Net: Learning Dense Volumetric Segmentation from Sparse Annotation. International Conference on Medical Image Computing and Computer-Assisted Intervention (2016).
47. Vaswani, A. et al. Attention is All you Need. Adv. Neural Inf. Process. Syst. (2017).
48. Dosovitskiy, A. et al. An Image is Worth
49. Han, K. et al. A survey on Vision Transformer. IEEE Trans. Pattern Anal. Mach. Intell. 45, 87-110 (2023).
50. Liu, Z. et al. Swin Transformer: Hierarchical Vision Transformer using Shifted Windows. 2021 IEEE/CVF International Conference on Computer Vision (ICCV) 9992-10002 (2021).
51. Hatamizadeh, A., Yang, D., Roth, H. R. & Xu, D. UNETR: Transformers for 3D Medical Image Segmentation. 2022 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV). 1748-1758 (2021).
© The Author(s) 2024
Beijing Yakebot Technology Co., Ltd., Beijing, China; School of Mechanical Engineering and Automation, Beihang University, Beijing, China; State Key Laboratory of Oral & Maxillofacial Reconstruction and Regeneration, National Clinical Research Center for Oral Diseases, Shaanxi Key Laboratory of Stomatology, Digital Center, School of Stomatology, The Fourth Military Medical University, Xi’an, China and Key Laboratory of Biomechanics and Mechanobiology of the Ministry of Education, Beijing Advanced Innovation Center for Biomedical Engineering, School of Biological Science and Medical Engineering, Beihang University, Beijing, China
Correspondence: Yimin Zhao(xaddoor@foxmail.com)
These authors contributed equally: Yu Liu, Rui Xie
DOI: https://doi.org/10.1038/s41368-024-00294-z
PMID: https://pubmed.ncbi.nlm.nih.gov/38719817
Publication Date: 2024-05-08
Fully automatic AI segmentation of oral surgery-related tissues based on cone beam computed tomography images
Abstract
Accurate segmentation of oral surgery-related tissues from cone beam computed tomography (CBCT) images can significantly accelerate treatment planning and improve surgical accuracy. In this paper, we propose a fully automated tissue segmentation system for dental implant surgery. Specifically, we propose an image preprocessing method based on data distribution histograms, which can adaptively process CBCT images with different parameters. Based on this, we use the bone segmentation network to obtain the segmentation results of alveolar bone, teeth, and maxillary sinus. We use the tooth and mandibular regions as the ROI regions of tooth segmentation and mandibular nerve tube segmentation to achieve the corresponding tasks. The tooth segmentation results can obtain the order information of the dentition. The corresponding experimental results show that our method can achieve higher segmentation accuracy and efficiency compared to existing methods. Its average Dice scores on the tooth, alveolar bone, maxillary sinus, and mandibular canal segmentation tasks were
INTRODUCTION
accurate postoperative evaluation,

- Tooth segmentation results of existing methods do not mark the teeth according to the FDI Two-Digit Notation,
so that the missing tooth information cannot be accurately detected and represented. We reproduced the method of Cui et al. which is the state-of-the-art in this field, the result is shown in Fig. 1. It cannot achieve the same classification results for teeth in the same position, which is mapped in the figure as different colors of teeth in the same position. The method proposed by Liu et al. requires first segmenting the oral scan model and registering it with the CBCT image to obtain segmentation results. This limits the application of these methods in the analysis and diagnosis of dental defects, since information on missing teeth is essential for the correct localization and assessment of the patient’s oral health status. - Unlike conventional CT, different CBCT equipment manufacturers use different imaging devices and interpolation methods, and also dynamically select imaging parameters according to the patient’s condition when in use, so CBCT images will have different grayscale distribution and contrast, which makes it difficult to apply simple image
preprocessing methods to all CBCT images; and in the tooth instance segmentation task, most of the original images are soft tissue regions that are not useful for segmentation, resulting in an unbalanced distribution of data, which is not addressed by the current methods. The actual CBCT images are shown in Fig. 2, and it can be seen that there are large differences in image contrast, field of view, etc. - Existing methods are too complex and often require multiple steps to obtain good segmentation results. These methods may require the use of multiple preprocessing techniques, feature extraction methods, classifiers or models, and post-processing steps to complete the segmentation of teeth. This complexity leads to less computationally efficient methods, each step may introduce errors or mistakes, and each step in the whole process requires careful tuning and validation, which increases the difficulty of method development and application.
- Current methods do not perform all these segmentation tasks completely automatically in an end-to-end manner, as they usually focus on a single task, such as tooth segmentation or alveolar bone segmentation on predefined region of interest (ROI), with little research on the segmentation of the mandibular canal from the maxillary sinus.
RESULTS


| Segment classes | Dice/% | mloU/% | HD/mm | ASD/mm |
| Tooth |
|
|
|
|
| Maxillary bone |
|
|
|
|
| Mandible bone |
|
|
|
|
| Mandibular canal |
|
|
|
|
| Maxillary sinus |
|
|
|
|
| Average |
|
|
|
|

| Table 2. Experimental results of quantitative comparison with existing advanced methods in terms of segmentation and detection accuracy | ||||
| Methods | Dice/% | mloU/% | HD/mm | ASD/mm |
| Hi-MoToothSeg |
|
|
|
|
| nnUNet |
|
|
|
|
| ToothNet |
|
|
|
|
| RELU-Net |
|
|
|
|
| DenseASPP-UNet |
|
|
|
|
| Ours |
|
|
|
|
| Bold text represents the highest value in its column | ||||
| Table 3. Analysis of the effect of choosing different relationships between
|
||||
| Items | Dice/% | mloU/% | HD/mm | ASD/mm |
| Normal preprocess |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
| Bold text represents the highest value in its column | ||||
DISCUSSION

| Methods | T1 | T2 | T3 | T4 | T5 | T6 | T7 | T8 |
| Ours |
|
|
|
|
|
|
|
|
| Manufacturer | Manufacturer’s model name | Dice/% | mloU/% | HD/mm | ASD/mm |
| Carestream Health | CS 9300, CS 9301 |
|
|
|
|
| Imaging Sciences International | 9-17 |
|
|
|
|
| J.Morita.Mfg.Corp. |
|
|
|
|
|
| LargeV | HighRes3D, SMART3D |
|
|
|
|
| NewTom | NTVGiMK4, NTVGiEVO, NT5G |
|
|
|
|
| NNT | NTVGiEVO |
|
|
|
|
| PaloDEx Group Oy | ORTHOPANTOMOGRAPH OP 3D |
|
|
|
|
| RAY Co., Ltd. | RAYSCAN N Alpha Plus |
|
|
|
|
| Sirona | ORTHOPHOS SL |
|
|
|
|
| Vatech Company Limited | PHT-35LHS |
|
|
|
|
| YOFO | Pirox-R |
|
|
|
|
| Items | Al | Expert | Al-assist + hand-tuning |
| Time
|
|
240 | 5.6 |
| Dice/% |
|
92.6 | |
| Bold text represents the highest value in its column | |||
algorithm difficult and make it impossible to accurately locate and select specific teeth during software interaction. The ability to accurately capture information about missing teeth makes our method more conducive to comprehensive dental evaluation.
CONCLUSION
METHODS
CBCT image adaptive preprocessing method
Network structure

it difficult to obtain the relationship between two teeth that are far away when performing tooth instance segmentation, which leads to low segmentation accuracy. The existing CNN-based tooth instance segmentation methods require other feature information as input, such as pre-obtained tooth centroids, tooth positions obtained using oral scan models, etc.


simultaneously model the relationships between long-distance pixels and extract local information, which is crucial for predicting tooth position.
Swin-Transformer: as a feature extractor, used to extract meaningful feature representations from the input image. It is based on the Swin-Transformer architecture
| Embed dimension | Feature size | Number of blocks | Window size | Number of heads | Parameters | FLOPS |
| 768 | 48 |
|
|
|
62.19 M | 394.84 G |
| Manufacturer | Manufacturer’s model name | Sex (Female/ Male) | Tube voltage/kVp | Tube current
|
Spacing/ mm | Average age/ years | CBCT number (cases) |
| Carestream Health | CS 9300, CS 9301 | 90 | 10 | 0.18 | 40.1 | 15 | |
| International | |||||||
| J.Morita.Mfg.Corp. | 2F/2M | 89 | 7 | 0.25 | 25.3 | 4 | |
| LargeV | HighRes3D, SMART3D |
|
100 | 4 | 0.25 | 44.5 | 112 |
| NewTom | NTVGiMK4, NTVGiEVO, NT5G | 22F/13M | 110 | 1, 2, 3, 4, 5, 7, 9, 10 | 0.3, 0.25 | 42.1 | 35 |
| NNT | NTVGiEVO | 7M | 110 | 3, 7, 8, 9, 10, 11, 14 | 0.3 | 31.2 | 7 |
| PaloDEx Group Oy | ORTHOPANTOMOGRAPH OP 3D | 5M | 95 | 3, 8 | 0.25 | 44.2 | 7 |
| RAY Co., Ltd. | RAYSCAN N Alpha Plus | 1F/2M | 90 | 10 | 1 | 3 | |
| Sirona | ORTHOPHOS SL | 85 | 10 | 0.22 | 46.7 | 10 | |
| Vatech Company Limited | PHT-35LHS | 1F/3M | 94 | 8 | 0.2 | 47.8 | 4 |
| YOFO | Pirox-R | 29F/23M | 90 | 8 | 0.25 | 40.1 | 52 |
| 0.4 | 91 | ||||||
| 0.4 | 97 | ||||||
Loss function
respectively, which are both set to 1 in this experiment.
Dataset
Evaluation metrics
Implementation details
ACKNOWLEDGEMENTS
AUTHOR CONTRIBUTIONS
ADDITIONAL INFORMATION
REFERENCES
- Bai, S. Z. et al. [Animal experiment on the accuracy of the Autonomous Dental Implant Robotic System]. Zhonghua Kou Qiang Yi Xue Za Zhi 56, 170-174 (2021).
- Jia, S., Wang, G., Zhao, Y. & Wang, X. Accuracy of an autonomous dental implant robotic system versus static guide-assisted implant surgery: a retrospective clinical study. J. Prosthet. Dent. https://doi.org/10.1016/j.prosdent.2023.04.027 (2023).
- Li, Z., Xie, R., Bai, S. & Zhao, Y. Implant placement with an autonomous dental implant robot: a clinical report. J. Prosthet. Dent. https://doi.org/10.1016/ j.prosdent.2023.02.014 (2023).
- Wu, Q., Zhao, Y. M., Bai, S. Z. & Li, X. Application of robotics in stomatology. Int. J. Comput. Dent. 22, 251-260 (2019).
- Cheng, K. J. et al. Accuracy of dental implant surgery with robotic position feedback and registration algorithm: an in-vitro study. Comput. Biol. Med. 129, 104153 (2021).
- Tao, B. et al. Accuracy of dental implant surgery using dynamic navigation and robotic systems: an in vitro study. J. Dent. 123, 104170 (2022).
- Bolding, S. L. & Reebye, U. N. Accuracy of haptic robotic guidance of dental implant surgery for completely edentulous arches. J. Prosthet. Dent. 128, 639-647 (2022).
- Yang, X. et al. ImplantFormer: vision transformer based implant position regression using dental CBCT data. Accessed July 3, 2023. Preprint at http:// arxiv.org/abs/2210.16467 (2023).
- Płotka, S. et al. Convolutional neural networks in orthodontics: a review. Accessed July 3, 2023. Preprint at http://arxiv.org/abs/2104.08886 (2021).
- Duy, N. T., Lamecker, H., Kainmueller, D. & Zachow, S. Automatic detection and classification of teeth in CT data. In Medical Image Computing and ComputerAssisted Intervention – MICCAI 2012, Lecture Notes in Computer Science (eds Ayache, N., Delingette, H., Golland, P. & Mori, K.) 609-616 (Springer, 2012).
- Barone, S., Paoli, A. & Razionale, A. V. CT segmentation of dental shapes by anatomy-driven reformation imaging and B-spline modelling. Int. J. Numer. Method. Biomed. Eng. 32, e02747 (2016).
- Sepehrian, M., Deylami, A. M. & Zoroofi, R. A. Individual teeth segmentation in CBCT and MSCT dental images using watershed. In 2013 20th Iranian Conference on Biomedical Engineering (ICBME) 27-30 (IEEE, 2013).
- Gao, H. & Chae, O. Individual tooth segmentation from CT images using level set method with shape and intensity prior. Pattern Recognit. 43, 2406-2417 (2010).
- Qian, J. et al. An automatic tooth reconstruction method based on multimodal data. J. Vis. 24, 205-221 (2021).
- Poonsri, A., Aimjirakul, N., Charoenpong, T. & Sukjamsri, C. Teeth segmentation from dental X-ray image by template matching. In 2016 9th Biomedical Engineering International Conference (BMEICON) 1-4 (IEEE, 2016).
- Akhoondali, H., Zoroofi, R. A. & Shirani, G. Rapid automatic segmentation and visualization of teeth in CT-scan data. J. Appl. Sci. 9, 2031-2044 (2009).
- Zou, X., Liu, W., Wang, J., Tao, R. & Zheng, G. ARST: auto-regressive surgical transformer for phase recognition from laparoscopic videos. Accessed July 3, 2023. Preprint at http://arxiv.org/abs/2209.01148 (2022).
- Huang, S., Xu, T., Shen, N., Mu, F. & Li, J. Rethinking few-shot medical segmentation: a vector quantization view.
- Zhang, J., Xie, Y., Xia, Y. & Shen, C. DoDNet: learning to segment multi-organ and tumors from multiple partially labeled datasets. Accessed July 3, 2023. Preprint at http://arxiv.org/abs/2011.10217 (2020).
- Wang, R. et al. Medical image segmentation using deep learning: a survey. IET Image Process. 16, 1243-1267 (2022).
- Zhang, R. et al. Multiple supervised residual network for osteosarcoma segmentation in CT images. Comput. Med. Imaging Graph. 63, 1-8 (2018).
- Tao, R., Liu, W. & Zheng, G. Spine-transformers: vertebra labeling and segmentation in arbitrary field-of-view spine CTs via 3D transformers. Med. Image Anal. 75, 102258 (2022).
- Cai, Y. et al. Swin Unet3D: a three-dimensional medical image segmentation network combining vision transformer and convolution. BMC Med. Inf. Decis. Mak. 23, 33 (2023).
- Cao, H. et al. Swin-Unet: Unet-Like Pure Transformer for Medical Image Segmentation. In Computer Vision – ECCV 2022 Workshops. ECCV 2022. Lecture Notes in Computer Science, Vol. 13803 (eds. Karlinsky, L., Michaeli, T. & Nishino, K.) https://doi.org/10.1007/978-3-031-25066-8_9 (Springer, Cham, 2023).
12
25. Hatamizadeh, A. et al. Swin UNETR: Swin Transformers for Semantic Segmentation of Brain Tumors in MRI Images. https://arxiv.org/abs/2201.01266 (2022).
26. Milletari, F., Navab, N. & Ahmadi, S.-A. V-Net: Fully Convolutional Neural Networks for Volumetric Medical Image Segmentation. 2016 Fourth International Conference on 3D Vision (3DV), Stanford, CA, USA. 565-571, https://doi.org/10.1109/3DV.2016.79 (2016).
27. Cui, Z. et al. A fully automatic AI system for tooth and alveolar bone segmentation from cone-beam CT images. Nat. Commun. 13, 2096 (2022).
28. Chen, Y. et al. Automatic segmentation of individual tooth in dental CBCT images from tooth surface map by a multi-task FCN. IEEE Access 8, 97296-97309 (2020).
29. Jaskari, J. et al. Deep learning method for mandibular canal segmentation in dental cone beam computed tomography volumes. Sci. Rep. 10, 5842 (2020).
30. Verhelst, P. J. et al. Layered deep learning for automatic mandibular segmentation in cone-beam computed tomography. J. Dent. 114, 103786 (2021).
31. Chung, M. et al. Pose-aware instance segmentation framework from cone beam CT images for tooth segmentation. Comput. Biol. Med. 120, 103720 (2020).
32. Cui, Z., Li, C. & Wang, W. ToothNet: automatic tooth instance segmentation and identification from cone beam CT Images. In 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 6361-6370 (IEEE, 2019).
33. Lee, J. et al. Tooth instance segmentation from cone-beam CT images through point-based detection and Gaussian disentanglement. Multimed Tools Appl. 81, 18327-18342 (2022).
34. Gerhardt, M. D. N. et al. Automated detection and labelling of teeth and small edentulous regions on cone-beam computed tomography using convolutional neural networks. J. Dent. 122, 104139 (2022).
35. Liu, J. et al. Deep learning-enabled 3D multimodal fusion of cone-beam CT and intraoral mesh scans for clinically applicable tooth-bone reconstruction. Patterns 4, 100825 (2023).
36. Nackaerts, O. et al. Segmentation of trabecular jaw bone on cone beam CT datasets: segmentation of jaw bone on CBCT datasets. Clin. Implant Dent. Relat. Res. 17, 1082-1091 (2015).
37. FDI World Dental Federation. Accessed July 11, 2023. https://web.archive.org/web/ 20070401074213/http://www.fdiworldental.org/resources/5_Onotation.html (2007).
38. Cui, Z., et al. Hierarchical morphology-guided tooth instance segmentation from CBCT images. In Information Processing in Medical Imaging, Lecture Notes in Computer Science (eds Feragen, A., Sommer, S., Schnabel, J. & Nielsen, M.) 150-162 (Springer International Publishing, 2021).
39. Isensee, F., Jäger, P. F., Kohl, S. A. A., Petersen, J. & Maier-Hein, K. H. Automated design of deep learning methods for biomedical image segmentation. Nat. Methods 18, 203-211 (2021).
40. Wu, X. et al. Center-sensitive and boundary-aware tooth instance segmentation and classification from cone-beam CT. In 2020 IEEE 17th International Symposium on Biomedical Imaging (ISBI) 939-942 (IEEE, 2020).
41. Zhang, Y. & Yu, H. Convolutional neural network based metal artifact reduction in X-ray computed tomography. IEEE Trans. Med. Imaging 37, 1370-1381 (2018).
42. Liang, X. et al. Generating synthesized computed tomography (CT) from conebeam computed tomography (CBCT) using CycleGAN for adaptive radiation therapy. Phys. Med. Biol. 64, 125002 (2019).
43. Liu, Y. et al. CBCT-based synthetic CT generation using deep-attention cycleGAN for pancreatic adaptive radiotherapy. Med. Phys. 47, 2472-2483 (2020).
44. Galibourg, A. et al. Assessment of automatic segmentation of teeth using a watershed-based method. Dentomaxillofac. Radiol. 47, 20170220 (2018).
45. Hounsfield, G. N. Computed medical imaging. Science 210, 22-28 (1980).
46. Özgün, Ç., Abdulkadir, A., Lienkamp, S. S., Brox, T. & Ronneberger, O. 3D U-Net: Learning Dense Volumetric Segmentation from Sparse Annotation. International Conference on Medical Image Computing and Computer-Assisted Intervention (2016).
47. Vaswani, A. et al. Attention is All you Need. Adv. Neural Inf. Process. Syst. (2017).
48. Dosovitskiy, A. et al. An Image is Worth
49. Han, K. et al. A survey on Vision Transformer. IEEE Trans. Pattern Anal. Mach. Intell. 45, 87-110 (2023).
50. Liu, Z. et al. Swin Transformer: Hierarchical Vision Transformer using Shifted Windows. 2021 IEEE/CVF International Conference on Computer Vision (ICCV) 9992-10002 (2021).
51. Hatamizadeh, A., Yang, D., Roth, H. R. & Xu, D. UNETR: Transformers for 3D Medical Image Segmentation. 2022 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV). 1748-1758 (2021).
© The Author(s) 2024
Beijing Yakebot Technology Co., Ltd., Beijing, China; School of Mechanical Engineering and Automation, Beihang University, Beijing, China; State Key Laboratory of Oral & Maxillofacial Reconstruction and Regeneration, National Clinical Research Center for Oral Diseases, Shaanxi Key Laboratory of Stomatology, Digital Center, School of Stomatology, The Fourth Military Medical University, Xi’an, China and Key Laboratory of Biomechanics and Mechanobiology of the Ministry of Education, Beijing Advanced Innovation Center for Biomedical Engineering, School of Biological Science and Medical Engineering, Beihang University, Beijing, China
Correspondence: Yimin Zhao(xaddoor@foxmail.com)
These authors contributed equally: Yu Liu, Rui Xie
