DOI: https://doi.org/10.1016/j.imavis.2024.105057
تاريخ النشر: 2024-05-01
ASF-YOLO: نموذج YOLO جديد مع دمج تسلسل المقياس الانتباهي لتجزئة حالات الخلايا
معلومات المقال
الكلمات المفتاحية:
تقسيم الأجسام الصغيرة
تنظر مرة واحدة فقط (YOLO)
دمج ميزات التسلسل
آلية الانتباه
الملخص
نقترح إطار عمل جديد يعتمد على دمج تسلسل المقياس الانتباهي (ASF-YOLO) والذي يجمع بين الميزات المكانية والقياسية لتحقيق تقسيم دقيق وسريع لعينات الخلايا. يعتمد هذا الإطار على إطار عمل تقسيم YOLO، حيث نستخدم وحدة دمج ميزات تسلسل المقياس (SSFF) لتعزيز قدرة الشبكة على استخراج المعلومات متعددة المقاييس، ووحدة تشفير الميزات الثلاثية (TFE) لدمج خرائط الميزات من مقاييس مختلفة لزيادة المعلومات التفصيلية. كما نقدم آلية انتباه القناة والموقع (CPAM) لدمج كل من وحدات SSFF وTFE، والتي تركز على القنوات المعلوماتية والأجسام الصغيرة المرتبطة بالموقع المكاني لتحسين أداء الكشف والتقسيم. تظهر التحقق التجريبي على مجموعتين من بيانات الخلايا دقة وسرعة تقسيم ملحوظة لنموذج ASF-YOLO المقترح. حيث يحقق معدل دقة الصندوق (box mAP) 0.91، ومعدل دقة القناع (mask mAP) 0.887، وسرعة استدلال تبلغ 47.3 إطارًا في الثانية على مجموعة بيانات 2018 Data Science Bowl، متفوقًا على أحدث الأساليب. الشيفرة المصدرية متاحة على https://github.com/mkang315/ASF-YOLO.
1. المقدمة
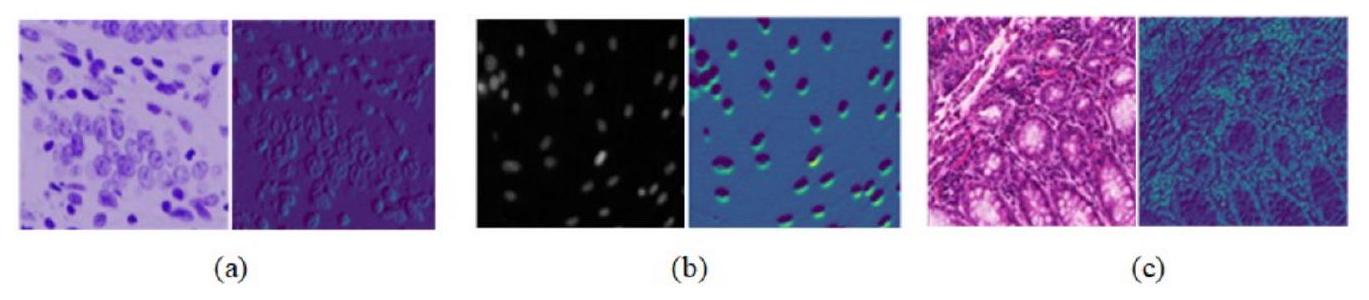
تؤدي الحدود غير الواضحة للخلايا إلى دقة تقسيم ضعيفة. يتطلب الأمر تقسيمًا دقيقًا ومفصلاً لأنواع مختلفة من الكائنات في صور الخلايا. كما هو موضح في الشكل 1، فإن أنواع صور الخلايا المختلفة لديها اختلافات كبيرة في اللون، والشكل، والملمس، ومعلومات أخرى مميزة بسبب اختلافات في شكل الخلايا، وطرق التحضير، وتقنيات التصوير. على الرغم من تحسين دقة وسرعة التقسيم للصور الطبيعية، يمكن تحسين هياكل نماذج YOLO بشكل أكبر للتعامل مع الكائنات الصغيرة في الصور الطبية، مثل الخلايا.


جزء من النموذج. تشمل هذه دمج الميزات متعددة المقاييس ودمج آلية الانتباه.
- نحن نصمم وحدة دمج ميزات تسلسل المقياس (SSFF) ووحدة ترميز الميزات الثلاثية (TFE) لدمج خرائط الميزات متعددة المقاييس المستخرجة من الهيكل العظمي في هيكل شبكة تجميع المسار (PANet) [16]. تجمع SSFF المعلومات الدلالية العالمية للصور عبر مقاييس مختلفة من خلال التطبيع، والتكبير، ودمج الميزات متعددة المقاييس في عملية تلافيف ثلاثية الأبعاد. وبالتالي، يمكنها التعامل بفعالية مع الأجسام ذات الأحجام، والاتجاهات، ونسب الأبعاد المتنوعة في تمثيل مساحة المقياس لتحسين تقسيم الأجسام. تتضمن TFE خرائط ميزات صغيرة ومتوسطة وكبيرة الحجم لالتقاط المعلومات المكانية الدقيقة للأجسام الصغيرة عبر مقاييس مميزة. تتغلب هذه على قيود FPN في YOLOv5، التي لا يمكنها استغلال العلاقات بين خرائط الميزات الهرمية بالكامل عبر عمليات الجمع البسيطة والدمج، وتعتمد بشكل أساسي على خرائط الميزات الصغيرة.
- ثم نصمم آلية انتباه القناة والموقع (CPAM) لدمج معلومات الميزات من وحدات SSFF وTFE. تتيح هذه الوحدة للنموذج ضبط تركيزه بشكل تكيفي على القنوات والمواقع المكانية ذات الصلة بالأجسام الصغيرة عبر مقاييس مختلفة، وبالتالي تحقيق تقسيم أفضل للحالات مقارنة بهيكل YOLOv5 التقليدي بدون آلية انتباه.
- نطبق نموذج ASF-YOLO المقترح لمهام تقسيم الحالات الصعبة للأجسام المتداخلة بكثافة وأنواع خلايا متنوعة. حسب علمنا، هذه هي أول دراسة تستفيد من نموذج قائم على YOLO لتقسيم حالات الخلايا. تُظهر تقييمات مجموعتي بيانات الخلايا المرجعية دقة اكتشاف وسرعة متفوقة مقارنة بأساليب أخرى متطورة، بما في ذلك النماذج المعتمدة على CNN التي تم استخدامها سابقًا لتقسيم الخلايا والعديد من النماذج الحديثة المعتمدة على YOLO.
2. الأعمال ذات الصلة
2.1. تقسيم حالات الخلايا
2.2. تحسين YOLO لتقسيم الحالات
[38] قناع الحالة بكفاءة من خلال الانحدار المباشر والواضح للحدود باستخدام قيود متعددة الطلبات مصممة تتكون من خسارة المسافة القطبية وخسارة القطاع. بالإضافة إلى ذلك، دمجت النماذج الهجينة YOLOMask [39] وYUSEG [40] YOLOv4 المحسن [12] وYOLOv5s الأصلي مع شبكة تقسيم دلالي U-Net لضمان دقة تقسيم الحالات. حسب علمنا، لم يتم تطبيق هذه الهياكل المحسنة لـ YOLO، التي تم تصميمها في الأصل لتقسيم الحالات للصور الطبيعية، على تقسيم حالات الخلايا، والتي تكون أكثر تحديًا بسبب الخلايا الصغيرة والمتداخلة بكثافة.
3. نموذج ASF-YOLO المقترح
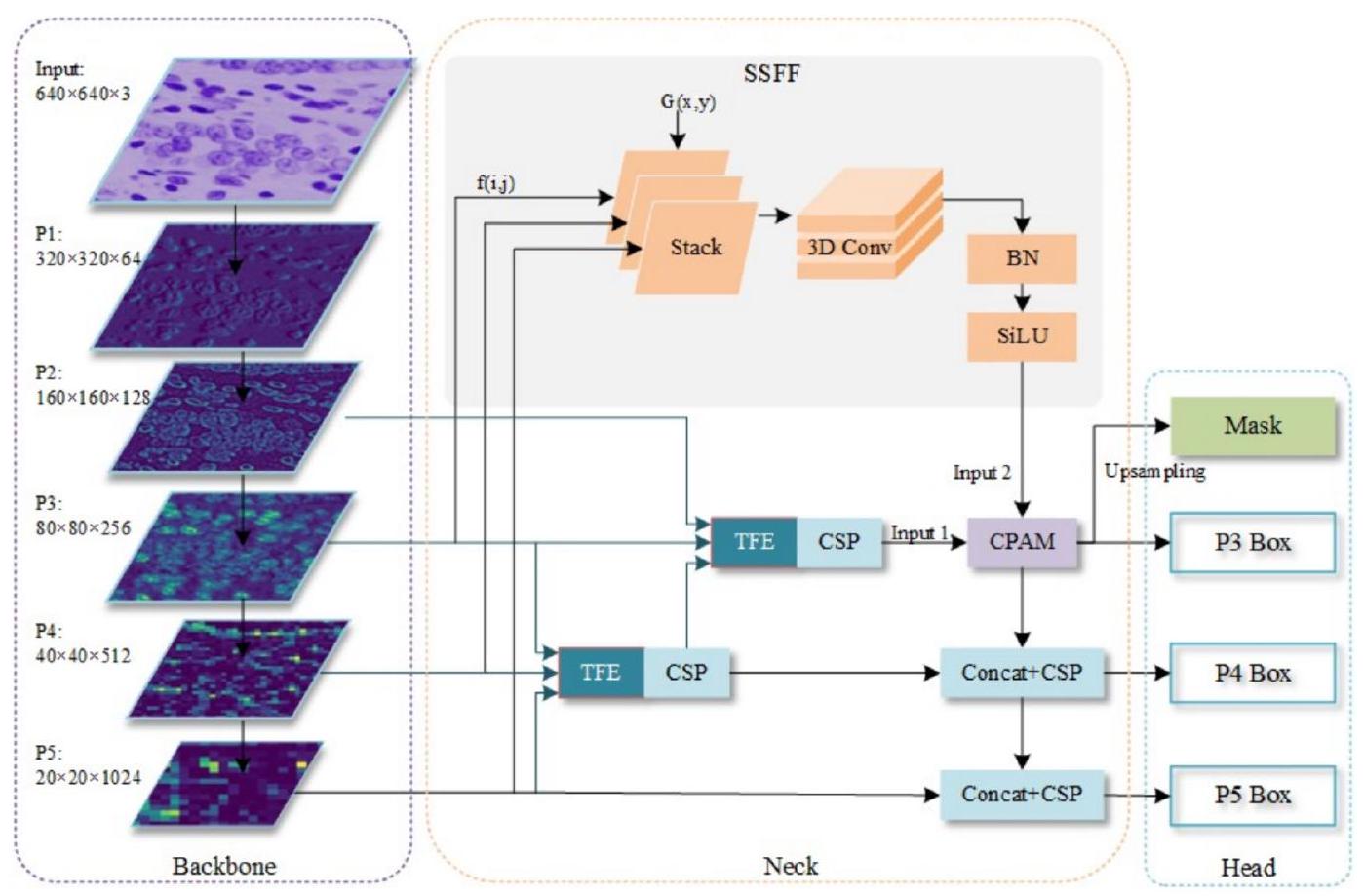
3.1. الهيكل العام
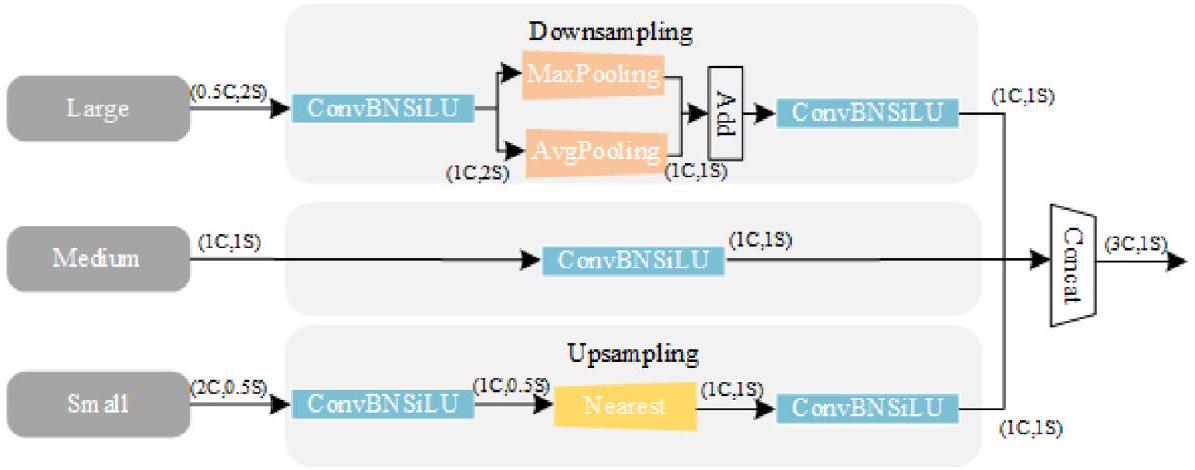
3.2. وحدة دمج ميزات تسلسل المقياس
- أ
يتم استخدام الالتفاف لتغيير عدد القنوات في مستويات الميزات P4 و P5 إلى 256. - تُستخدم طريقة الاستيفاء بأقرب جار [45] لضبط حجمها ليتناسب مع حجم مستوى P3.
- تُستخدم طريقة “unsqueeze” لزيادة أبعاد كل طبقة ميزات، مما يغيرها من موتر ثلاثي الأبعاد [الارتفاع، العرض، القناة] إلى موتر رباعي الأبعاد [العمق، الارتفاع، العرض، القناة].
- ثم يتم دمج خرائط الميزات ذات الأبعاد الأربعة على طول بعد العمق لتشكيل خريطة ميزات ثلاثية الأبعاد للت convolutions اللاحقة.
- أخيرًا، يتم استخدام الالتفاف ثلاثي الأبعاد، وتطبيع الدفعات ثلاثي الأبعاد، ودالة التنشيط SiLU [46] لإكمال استخراج ميزات تسلسل المقياس.
3.3. وحدة ترميز الميزة الثلاثية
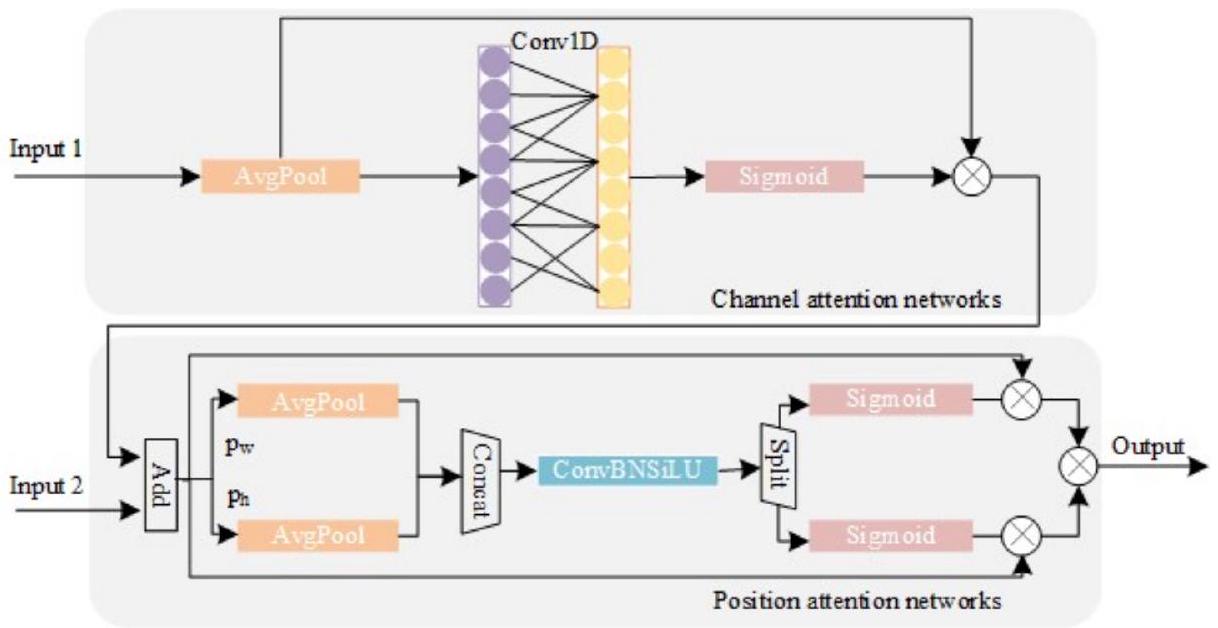
3.4. آلية الانتباه للقناة والموقع
ميزات القناة.
دمج مخرجات آلية انتباه القناة مع الميزات من SSFF (الإدخال 2) كمدخل لشبكة انتباه الموقع يوفر معلومات تكميلية لاستخراج معلومات الموقع الحيوية من كل خلية. على عكس آلية انتباه القناة، تقوم آلية انتباه الموقع أولاً بتقسيم خريطة الميزات المدخلة إلى جزئين من حيث العرض والارتفاع، والتي تتم معالجتها بشكل منفصل لترميز الميزات في المحاور.
3.5. تحسين صناديق التثبيت
أين
4. التجارب
4.1. مجموعات البيانات
مقارنة أداء النماذج المختلفة لتجزئة مثيلات الخلايا على مجموعة بيانات DSB2018. أفضل النتائج بالخط العريض.
| نموذج | بارام (م) | صندوق | قناع | |||
|
|
|
|
|
FPS | ||
| ماسک R-CNN [2] | ٤٣.٧٥ | 0.774 | 0.519 | 0.782 | 0.525 | 20 |
| قناع كاسكيد RCNN [3] | 69.17 | – | – | 0.783 | 0.533 | 17.9 |
| سولو [4] | – | – | – | 0.642 | 0.398 | ٢٥ |
| SOLOv2 [5] | – | – | – | 0.741 | 0.495 | ٢٨.٧ |
| YOLACT [6] | ٣٤.٣ | 0.703 | 0.456 | 0.683 | 0.440 | ٢٥ |
| ماسک RCNN سويين T [52] | ٤٧.٣ | 0.784 | 0.524 | 0.783 | 0.527 | ٢٤ |
| YOLOv5l-seg [8] | ٤٥.٢٧ | 0.876 | 0.616 | 0.855 | 0.502 | ٤٦.٩ |
| YOLOv8l-seg [9] | ٤٥.٩١ | 0.865 | 0.631 | 0.866 | 0.562 | ٤٥.٥ |
| ASF-YOLO (خاصتنا) | ٤٦.١٨ | 0.910 | 0.676 | 0.887 | 0.558 | ٤٧.٣ |
مقارنة أداء النماذج المختلفة لتجزئة مثيلات الخلايا على مجموعة بيانات BCC. أفضل النتائج بالخط العريض.
| نموذج | صندوق | قناع | ||
|
|
|
|
|
|
| ماسک R-CNN [2] | 0.852 | 0.614 | 0.836 | 0.628 |
| كاسكيد ماسك آر-سي إن إن [3] | 0.836 | 0.630 | 0.823 | 0.598 |
| سولو [4] | – | – | 0.864 | 0.647 |
| SOLOv2 [5] | – | – | 0.860 | 0.651 |
| YOLACT [6] | 0.715 | 0.545 | 0.774 | 0.565 |
| ماسک RCNN سويين T [52] | 0.841 | 0.604 | 0.806 | 0.588 |
| YOLOv5l-seg [8] | 0.892 | 0.703 | 0.877 | 0.672 |
| YOLOv8l-seg [9] | 0.850 | 0.619 | 0.814 | 0.564 |
| ASF-YOLO (خاصتنا) | 0.911 | 0.737 | 0.898 | 0.645 |
4.2. تفاصيل التنفيذ
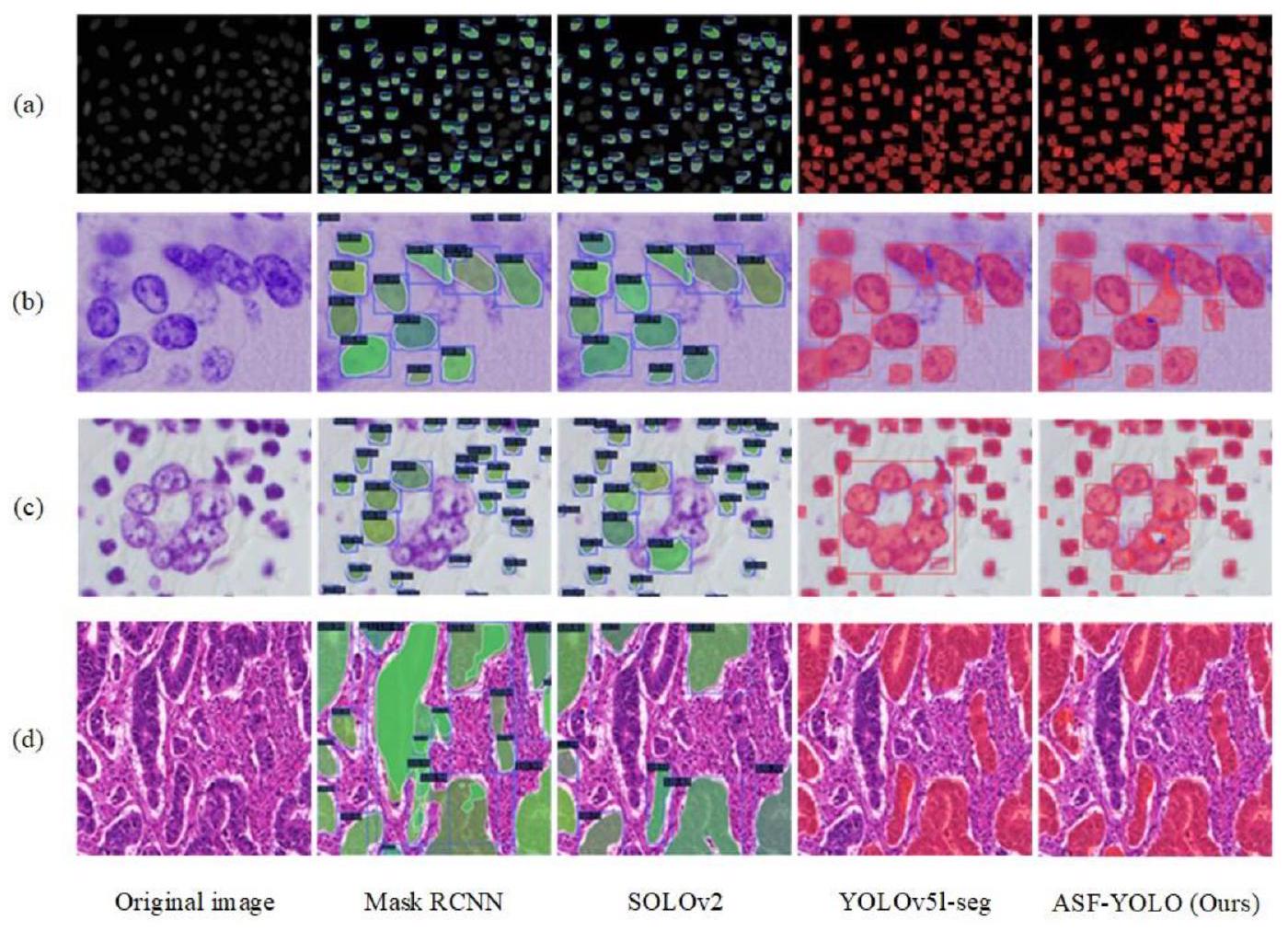
4.3. النتائج الكمية
دراسة الإزالة للمكونات الرئيسية لنموذج ASF-YOLO على مجموعة بيانات DSB2018.
| طريقة | صندوق | قناع | ||||||
| نظام عدم الحد من الصلابة | EIoU | تيفي | SSFF | CPAM |
|
|
|
|
|
|
0.876 | 0.616 | 0.855 | 0.502 | ||||
| 0.881 | 0.622 | 0.856 | 0.523 | |||||
|
|
0.880 | 0.634 | 0.852 | 0.507 | ||||
|
|
|
|
0.891 | 0.653 | 0.867 | 0.542 | ||
|
|
|
|
0.896 | 0.653 | 0.876 | 0.549 | ||
|
|
|
|
|
0.902 | 0.656 | 0.874 | 0.543 | |
|
|
|
|
|
|
0.910 | 0.676 | 0.887 | 0.558 |
4.4. النتائج النوعية
4.5. دراسة الاستئصال
أثر آليات الانتباه المختلفة.
| طريقة |
|
بارام (ك) | عمليات النقطة العائمة في الثانية (م) | |
| ASF-YOLO بدون انتباه (الخط الأساسي) | 0.902 | 0.874 | 0 | 0 |
| +SENet [29] | 0.901 | 0.879 | +8.19 | +1.65 |
| + CBAM [35] | 0.905 | 0.884 | +16.48 | +3.94 |
| + CA [53] | 0.903 | 0.8881 | +12.32 | +1.32 |
| + CPAM (خاصتنا) | 0.910 | 0.888 | +12.23 | +2.96 |
4.5.1. تأثير الطرق المقترحة
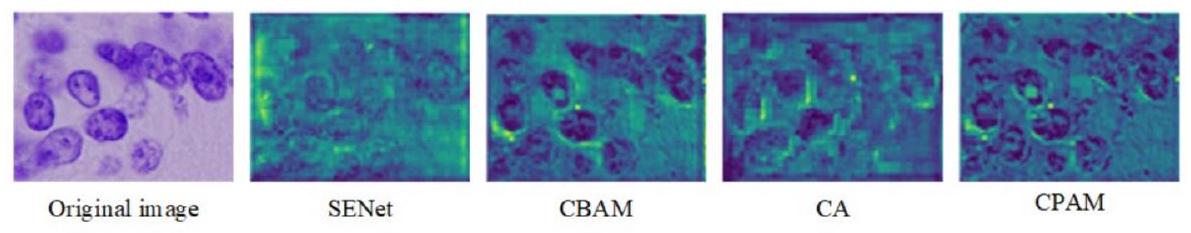
4.5.2. تأثير آليات الانتباه
تأثير وحدات الالتفاف المختلفة في العمود الفقري لنموذج ASFYOLO المقترح. أفضل النتائج بالخط العريض.
| مجموعة البيانات | الوحدة | الصندوق | القناع | ||
|
|
|
|
|
||
| DSB2018 | C3 | 0.910 | 0.676 | 0.887 | 0.558 |
| C2f | 0.867 | 0.633 | 0.859 | 0.558 | |
| BCC | C3 | 0.911 | 0.737 | 0.898 | 0.645 |
| C2f | 0.855 | 0.619 | 0.835 | 0.570 | |
4.5.3. تأثير وحدة الالتفاف في العمود الفقري
5. الخاتمة
لتحسين أداء الشبكات العصبية التلافيفية في التقاط المعلومات السياقية العالمية دون زيادة الجهد الحسابي وعدد المعلمات. كما يمكن إجراء التعلم بالنقل على شبكة العمود الفقري لاستخراج الميزات النسيجية، كما هو مستوحى من [56]. يمكن أيضًا اعتماد التقدم الأخير في Transformer لتحسين آلية الانتباه للنموذج المقترح.
بيان مساهمة المؤلفين
إعلان عن تضارب المصالح
توفر البيانات
الشكر والتقدير
References
[2] K. He, G. Gkioxari, P. Dollár, R. Girshick, Mask R-CNN, in: Proceedings of the IEEE/ CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2017, pp. 2980-2988.
[3] Z. Cai, N. Vasconcelos, Cascade R-CNN: delving into high quality object detection, in: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2018, pp. 6154-6162.
[4] X. Wang, T. Kong, C. Shen, Y. Jiang, L. Li, SOLO: segmenting objects by locations, in: A. Vedaldi, H. Bischof, T. Brox, J.-M. Frahm (Eds.), Computer Vision – ECCV 2020, Part XVIII, 2020, pp. 649-665.
[5] X. Wang, R. Zhang, T. Kong, L. Li, C. Shen, SOLOv2: dynamic and fast instance segmentation, in: H. Larochelle, M. Ranzato, R. Hadsell, M.F. Balcan, H. Lin (Eds.), Advances in Neural Information Processing Systems (NeurIPS), 2020, pp. 17721-17732.
[6] D. Bolya, C. Zhou, F. Xiao, Y.J. Lee, YOLACT: real-time instance segmentation, in: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2019, pp. 9158-9166.
[7] E. Mohamed, A. Shaker, A. El-Sallab, M. Hadhoud, INSTA-YOLO: real-time instance segmentation, arXiv:2102.0677, 2021.
[8] G. Jocher, YOLO by Ultralytics (version 5.7.0), GitHub, 2022. https://github.com/ ultralytics/yolov5.
[9] G. Jocher, A. Chaurasia, J. Qiu, YOLO by Ultralytics (version 8.0.0), GitHub, 2023. https://github.com/ultralytics/ultralytics.
[10] C.-Y. Wang, H.-Y.M. Liao, Y.-H. Wu, P.-Y. Chen, J.-W. Hsieh, I.-H. Yeh, CSPNet: A new backbone that can enhance learning capability of CNN, in: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), 2020, pp. 1571-1580.
[11] Ultralytics, Ultralytics YOLOv5 Architecture, Ultralytics, 2023. https://docs.ultr alytics.com/yolov5/tutorials/architecture_description.
[12] A. Bochkovskiy, C.Y. Wang, H.Y.M. Liao, YOLOv4: Optimal speed and accuracy of object detection, arXiv:2004.10934, 2020.
[13] T.-Y. Lin, P. Dollár, R. Girshick, K. He, B. Hariharan, S. Belongie, Feature pyramid networks for object detection, in: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2017, pp. 2117-2125.
[14] Y.-F. Zhang, W. Ren, Z. Zhang, Z. Jia, L. Wang, T. Tan, Focal and efficient IOU loss for accurate bounding box regression, Neurocomputing 506 (2022) 146-157.
[15] N. Bodla, B. Singh, R. Chellappa, L.S. Davis, Soft-NMS-improving object detection with one line of code, in: Proceedings of the IEEE International Conference on Computer Vision (ICCV), 2017, pp. 5562-5570.
[16] S. Liu, L. Qi, H. Qin, J. Shi, J. Jia, Path aggregation network for instance segmentation, in: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2018, pp. 8759-8768.
[17] R. Hollandi, N. Moshkov, L. Paavolainen, E. Tasnadi, F. Piccinini, P. Horvath, Nucleus segmentation: towards automated solutions, Trends Cell Biol. 32 (4) (2021) 295-310.
[18] J. Johnson, Adapting mask-RCNN for automatic nucleus segmentation, arXiv: 1805.00500, 2018.
[19] H. Jung, B. Lodhi, J. Kang, An automatic nuclei segmentation method based on deep convolutional neural networks for histopathology images, BMC Biomed. Eng. 1 (2019) 24.
[20] S. Fujita, X.-H. Han, Cell detection and segmentation in microscopy images with improved mask R-CNN, in: I. Sato, B. Han (Eds.), Computer Vision – ACCV 2020 Workshops, 2020, pp. 58-70.
[21] B. Bancher, A. Mahbod, I. Ellinger, Improving mask R-CNN for nuclei instance segmentation in hematoxylin & eosin-stained histological images, in: M. Atzori, N. Burlutskiy, F. Ciompi, Z. Li, F. Minhas, H. Müller, et al. (Eds.), Proceedings of the MICCAI Workshop on Computational Pathology, PMLR 156, 2024, pp. 20-35.
[22] J. Yi, P. Wu, M. Jiang, Q. Huang, D.J. Hoeppner, D.N. Metaxas, Attentive neural cell instance segmentation, Med. Image Anal. 55 (2019) 228-240.
[23] Z. Cheng, A. Qu, A fast and accurate algorithm for nuclei instance segmentation in microscopy images, IEEE Access 8 (2020) 158679-158689.
[24] W. Liu, D. Anguelov, D. Erhan, C. Szegedy, S. Reed, C. Fu, et al., SSD: Single shot multibox detector, in: B. Leibe, J. Matas, N. Sebe, M. Welling (Eds.), Computer Vision – ECCV 2016, Part I, 2016, pp. 21-37.
[25] A. Mahbod, G. Schaefer, G. Dorffner, S. Hatamikia, R. Ecker, I. Ellinger, A dual decoder U-net-based model for nuclei instance segmentation in hematoxylin and eosin-stained histological images, Front. Med. 9 (2022) 978146.
[26] O. Ronneberger, P. Fischer, T. Brox, U-Net: Convolutional networks for biomedical image segmentation, in: N. Navab, J. Hornegger, W.M. Wells, A.F. Frangi (Eds.), Medical Image Computing and Computer-Assisted Intervention – MICCAI, Part III, 2015, pp. 234-241.
[27] T. Konopczyński, R. Heiman, P. Woźnicki, P. Gniewek, M.-C. Duvernoy, O. Hallatschek, et al., Instance segmentation of densely packed cells using a hybrid model of U-Net and mask R-CNN, in: L. Rutkowski, R. Scherer, M. Korytkowski, W. Pedrycz, R. Tadeusiewicz, J.M. Zurada (Eds.), Artificial Intelligence and Soft Computing (ICAISC), Part I, 2020, pp. 626-635.
[28] J. Wang, Z. Zhang, M. Wu, Y. Ye, S. Wang, Y. Cao, et al., Improved BlendMask: nuclei instance segmentation for medical microscopy images, IET Image Process. 17 (7) (2023) 2284-2296.
[29] J. Hu, L. Shen, G. Sun, Squeeze-and-excitation networks, in: Proceedings of IEEE/ CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2018, pp. 7132-7141.
[30] Z. Shang, X. Wang, Y. Jiang, Z. Li, J. Ning, Identifying rumen protozoa in microscopic images of ruminant with improved YOLACT instance segmentation, Biosyst. Eng. 215 (2022) 156-169.
[31] Y. Wang, Z. Ouyang, R. Han, Z. Yin, Z. Yang, YOLOMask: Real-time instance segmentation with integrating YOLOv5 and OrienMask, in: Proceedings of the IEEE 22nd International Conference on Communication Technology (ICCT), 2022, pp. 1646-1650.
[32] W. Yang, S. Chen, G. Chen, Q. Shi, PR-YOLO: Improved YOLO for fast protozoa classification and segmentation, Res. Square Preprint (2024), https://doi.org/ 10.21203/rs.3.rs-3199595/v1.
[33] X. Cao, Y. Su, X. Geng, Y. Wang, YOLO-SF: YOLO for fire segmentation detection, IEEE Access 11 (2023) 111079-111092.
[34] C.-Y. Wang, A. Bochkovskiy, H.-Y.M. Liao, YOLOv7: Trainable bag-of-freebies sets new state-of-the-art for real-time object detectors, in: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2023, pp. 7464-7475.
[35] S. Woo, J. Park, J.-Y. Lee, I.S. Kweon, CBAM: Convolutional block attention module, in: V. Ferrari, M. Hebert, C. Sminchisescu, Y. Weiss (Eds.), Computer Vision – ECCV 2018, Part VII, 2018, pp. 3-19.
[36] O.M. Lawal, YOLOv5-LiNet: a lightweight network for fruits instance segmentation, PLoS One 18 (3) (2023) e0282297.
[37] M. Yasir, L. Zhan, S. Liu, J. Wan, M.S. Hossain, A.T. Isiacik Çolak, et al., Instance segmentation ship detection based on improved Yolov7 using complex background SAR images, Front. Mar. Sci. 10 (2023) 1113669.
[38] H. Liu, W. Xiong, Y. Zhang, YOLO-CORE: contour regression for efficient instance segmentation, Mach. Intell. Res. 20 (2023) 716-728.
[39] J. Hua, T. Hao, L. Zeng, G. Yu, YOLOMask, an instance segmentation algorithm based on complementary fusion network, Math 9 (15) (2021) 1766.
[40] B. Bai, J. Tian, T. Wang, S. Luo, S. Lyu, YUSEG: Yolo and Unet is all you need for cell instance segmentation, in: NeurIPS 2022 Weakly Supervised Cell Segmentation in Multi-modality High-Resolution Microscopy Images, 2022.
[41] T. Lindeberg, Scale-Space Theory in Computer Vision, Springer, Cham, 1994, pp. 10-11.
[42] D.J. Lowe, Distinctive image features from scale-invariant keypoints, Int. J. Comput. Vis. 60 (2004) 91-110.
[43] D. Tran, L. Bourdev, R. Fergus, L. Torresani, M. Paluri, Learning spatiotemporal features with 3D convolutional networks, in: Proceedings of IEEE International Conference on Computer Vision (ICCV), 2015, pp. 4489-4497.
[44] H.-J. Park, J.-W. Kang, B.-G. Kim, ssFPN: scale sequence (
[45] O. Rukundo, H. Cao, Nearest neighbor value interpolation, Int. J. Adv. Comput. Sci. Appl. 3 (4) (2012) 25-30.
[46] S. Elfwing, E. Uchibe, K. Doya, Sigmoid-weighted linear units for neural network function approximation in reinforcement learning, Neural Netw. 107 (2018) 3-11.
[47] H. Rezatofighi, N. Tsoi, J. Gwak, A. Sadeghian, I. Reid, S. Savarese, Generalized intersection over union: a metric and a loss for bounding box pegression, in: Proceedings of IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2019, pp. 658-666.
[48] Z. Zheng, P. Wang, W. Liu, J. Li, R. Ye, D. Ren, Distance-IoU loss: faster and better learning for bounding box regression, Proc. AAAI Conf. Artific. Intellig. 34 (07) (2020) 12993-13000.
[49] A. Neubeck, L. Van Gool, Efficient non-maximum suppression, in: Proceedings of the 18th International Conference on Pattern Recognition (ICPR), 2006, pp. 850-855.
[50] A. Goodman, A. Carpenter, E. Park, J. Lefman, J. BoozAllen, K. Thomas, et al., 2018 Data Science Bowl, Kaggle, 2018. https://kaggle.com/competitions/data-s cience-bowl-2018.
[51] CBI, Breast Cancer Cell, UCSB, 2008. https://bioimage.ucsb.edu/research/bio-seg mentation.
[52] OpenMMLab, Mask-rcnn_swin-t, GitHub, 2022. https://github.com/open-mm lab/mmdetection/tree/main/configs/swin.
[53] Q. Hou, D. Zhou, J. Feng, Coordinate attention for efficient mobile network design, in: Proceedings of IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2021, pp. 13708-13717.
[54] P. Wu, H. Li, N. Zeng, F. Li, FMD-Yolo: an efficient face mask detection method for COVID-19 prevention and control in public, Image Vis. Comput. 117 (2022) 104341.
[55] X. Guo, Z. Wang, P. Wu, Y. Li, F.E. Alsaadi, N. Zeng, ELTS-net: an enhanced liver tumor segmentation network with augmented receptive field and global contextual information, Comput. Biol. Med. 169 (2024) 107879.
[56] L. Wang, Y. Jiao, Y. Diao, N. Zeng, R. Yu, A novel approach combined transfer learning and deep learning to predict TMB from histology image, Pattern Recogn. Lett. 135 (2020) 244-248.
- *Corresponding author.
E-mail address:ting.cheeming@monash.edu(C.-M.Ting).
DOI: https://doi.org/10.1016/j.imavis.2024.105057
Publication Date: 2024-05-01
ASF-YOLO:A novel YOLO model with attentional scale sequence fusion for cell instance segmentation
ARTICLE INFO
Keywords:
Small object segmentation
You only look once(YOLO)
Sequence feature fusion
Attention mechanism
Abstract
We propose a novel Attentional Scale Sequence Fusion based You Only Look Once(YOLO)framework(ASF- YOLO)which combines spatial and scale features for accurate and fast cell instance segmentation.Built on the YOLO segmentation framework,we employ the Scale Sequence Feature Fusion(SSFF)module to enhance the multiscale information extraction capability of the network,and the Triple Feature Encoder(TFE)module to fuse feature maps of different scales to increase detailed information.We further introduce a Channel and Position Attention Mechanism(CPAM)to integrate both the SSFF and TFE modules,which focus on informative channels and spatial position-related small objects for improved detection and segmentation performance.Experimental validations on two cell datasets show remarkable segmentation accuracy and speed of the proposed ASF-YOLO model.It achieves a box mAP of 0.91 ,mask mAP of 0.887 ,and an inference speed of 47.3 FPS on the 2018 Data Science Bowl dataset,outperforming the state-of-the-art methods.The source code is available at https://github. com/mkang315/ASF-YOLO.
1.Introduction
as blurred boundaries of cells,which may result in poor segmentation accuracy.It requires accurate detailed segmentation of different types of objects in cell images.As shown in Fig.1,different types of cell images have large differences in color,morphology,texture,and other charac- teristic information due to differences in cell morphology,preparation methods,and imaging technologies.Despite its improved segmentation accuracy and speed for natural images,the architectures of YOLO-based models can be further optimized for handling small objects in medical images,such as cells.


part of the model. These include the fusion of multiscale features and the incorporation of attention mechanism.
- We design a Scale Sequence Feature Fusion (SSFF) module and a Triple Feature Encoder (TFE) module to fuse the multiscale feature maps extracted from the backbone in a Path Aggregation Network (PANet) [16] structure. The SSFF combines global semantic information of images across different scales by normalizing, upsampling, and concatenating multiscale features into a 3D convolution. Thus, it can effectively handle objects of varying sizes, orientations, and aspect ratios in a scale-space representation to improve object segmentation. The TFE incorporates small, medium, and large-sized feature maps to capture the fine spatial information of small objects across distinctive scales. These overcome the limitations of FPN in YOLOv5, which cannot fully exploit the correlations between the pyramid feature maps via simple sum and concatenation operations and mainly leverages small feature maps.
- We then design a Channel and Position Attention Mechanism (CPAM) to integrate feature information from the SSFF and TFE modules. This module allows the model to adaptively adjust its focus on relevant channels and spatial locations relevant for the small objects at different scales, and hence better instance segmentation than the conventional YOLOv5 architecture with no attention mechanism.
- We apply the proposed ASF-YOLO model for challenging instance segmentation tasks of densely overlapping and various cell types. To our best knowledge, this is the first work to leverage a YOLO-based model for cell instance segmentation. Evaluation of two benchmarking cell datasets shows superior detection accuracy and speed compared to other state-of-the-art methods, including CNN-based models previously used for cell segmentation and several recent YOLO-based models.

2. Related work
2.1. Cell instance segmentation
2.2. Improved YOLO for instance segmentation
[38] enhanced the mask of the instance efficiently by explicit and direct contour regression using a designed multi-order constraint consisting of a polar distance loss and a sector loss. In addition, the hybrid models YOLOMask [39] and YUSEG [40] combined optimized YOLOv4 [12] and the original YOLOv5s with semantic segmentation U-Net network to ensure the accuracy of instance segmentation. To our best knowledge, these improved YOLO architectures, originally designed for instance segmentation for natural images, have not been applied for cell instance segmentation, which is more challenging due to the small and densely overlapping cells.
3. The proposed ASF-YOLO model
3.1. Overall architecture

3.2. Scale sequence feature fusion module
- A
- Nearest neighbor interpolation method [45] is used to adjust their size to the size of the P3 level.
- The unsqueeze method is used to increase the dimension of each feature layer, changing it from a 3D tensor [height, width, channel] to a 4D tensor [depth, height, width, channel].
- The 4D feature maps are then concatenated along the depth dimension to form a 3D feature map for subsequent convolutions.
- Finally, 3D convolution, 3D batch normalization, and SiLU [46] activation function are used to complete scale sequence feature extraction.
3.3. Triple feature encoding module

3.4. Channel and position attention mechanism
channel features.
Combining the outputs of the channel attention mechanism with the features from SSFF (Input 2) as input to the position attention network provides complementary information to extract crucial location information from each cell. In contrast to the channel attention mechanism, the position attention mechanism first splits the input feature map into two parts in terms of its width and height, which are then processed separately for feature encoding in the axes (
3.5. Anchor box optimization
where
4. Experiments
4.1. Datasets
Performance comparison of different models for cell instance segmentation on the DSB2018 dataset. The best results are in bold.
| Model | Param (M) | Box | Mask | |||
|
|
|
|
|
FPS | ||
| Mask R-CNN [2] | 43.75 | 0.774 | 0.519 | 0.782 | 0.525 | 20 |
| Cascade Mask RCNN [3] | 69.17 | – | – | 0.783 | 0.533 | 17.9 |
| SOLO [4] | – | – | – | 0.642 | 0.398 | 25 |
| SOLOv2 [5] | – | – | – | 0.741 | 0.495 | 28.7 |
| YOLACT [6] | 34.3 | 0.703 | 0.456 | 0.683 | 0.440 | 25 |
| Mask RCNN Swin T [52] | 47.3 | 0.784 | 0.524 | 0.783 | 0.527 | 24 |
| YOLOv5l-seg [8] | 45.27 | 0.876 | 0.616 | 0.855 | 0.502 | 46.9 |
| YOLOv8l-seg [9] | 45.91 | 0.865 | 0.631 | 0.866 | 0.562 | 45.5 |
| ASF-YOLO (Ours) | 46.18 | 0.910 | 0.676 | 0.887 | 0.558 | 47.3 |
Performance comparison of different models for cell instance segmentation on the BCC dataset. The best results are in bold.
| Model | Box | Mask | ||
|
|
|
|
|
|
| Mask R-CNN [2] | 0.852 | 0.614 | 0.836 | 0.628 |
| Cascade Mask R-CNN [3] | 0.836 | 0.630 | 0.823 | 0.598 |
| SOLO [4] | – | – | 0.864 | 0.647 |
| SOLOv2 [5] | – | – | 0.860 | 0.651 |
| YOLACT [6] | 0.715 | 0.545 | 0.774 | 0.565 |
| Mask RCNN Swin T [52] | 0.841 | 0.604 | 0.806 | 0.588 |
| YOLOv5l-seg [8] | 0.892 | 0.703 | 0.877 | 0.672 |
| YOLOv8l-seg [9] | 0.850 | 0.619 | 0.814 | 0.564 |
| ASF-YOLO (Ours) | 0.911 | 0.737 | 0.898 | 0.645 |
4.2. Implementation details
4.3. Quantitative results

Ablation study of main components of the ASF-YOLO on the DSB2018 dataset.
| Method | Box | Mask | ||||||
| Soft-NMS | EIoU | TFE | SSFF | CPAM |
|
|
|
|
|
|
0.876 | 0.616 | 0.855 | 0.502 | ||||
| 0.881 | 0.622 | 0.856 | 0.523 | |||||
|
|
0.880 | 0.634 | 0.852 | 0.507 | ||||
|
|
|
|
0.891 | 0.653 | 0.867 | 0.542 | ||
|
|
|
|
0.896 | 0.653 | 0.876 | 0.549 | ||
|
|
|
|
|
0.902 | 0.656 | 0.874 | 0.543 | |
|
|
|
|
|
|
0.910 | 0.676 | 0.887 | 0.558 |
4.4. Qualitative results
4.5. Ablation study
Effect of different attention mechanisms.
| Method |
|
Param (K) | FLOPs (M) | |
| ASF-YOLO w/o attention (baseline) | 0.902 | 0.874 | 0 | 0 |
| +SENet [29] | 0.901 | 0.879 | +8.19 | +1.65 |
| + CBAM [35] | 0.905 | 0.884 | +16.48 | +3.94 |
| + CA [53] | 0.903 | 0.8881 | +12.32 | +1.32 |
| + CPAM (ours) | 0.910 | 0.888 | +12.23 | +2.96 |
4.5.1. Effect of the proposed methods
4.5.2. Effect of attention mechanisms

Effect of different convolution modules in the backbone of the proposed ASFYOLO. The best results are in bold.
| Dataset | Module | Box | Mask | ||
|
|
|
|
|
||
| DSB2018 | C3 | 0.910 | 0.676 | 0.887 | 0.558 |
| C2f | 0.867 | 0.633 | 0.859 | 0.558 | |
| BCC | C3 | 0.911 | 0.737 | 0.898 | 0.645 |
| C2f | 0.855 | 0.619 | 0.835 | 0.570 | |
4.5.3. Effect of convolution module in the backbone
5. Conclusion
be used to enhance the performance of CNNs in capturing global contextual information without increasing the computational effort and the number of parameters. As inspired by [56], transfer learning could be performed on the backbone network to extract histological features. The recent advances in Transformer could also be adopted to improve the attention mechanism of the proposed model.
CRediT authorship contribution statement
Declaration of competing interest
Data availability
Acknowledgments
References
[2] K. He, G. Gkioxari, P. Dollár, R. Girshick, Mask R-CNN, in: Proceedings of the IEEE/ CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2017, pp. 2980-2988.
[3] Z. Cai, N. Vasconcelos, Cascade R-CNN: delving into high quality object detection, in: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2018, pp. 6154-6162.
[4] X. Wang, T. Kong, C. Shen, Y. Jiang, L. Li, SOLO: segmenting objects by locations, in: A. Vedaldi, H. Bischof, T. Brox, J.-M. Frahm (Eds.), Computer Vision – ECCV 2020, Part XVIII, 2020, pp. 649-665.
[5] X. Wang, R. Zhang, T. Kong, L. Li, C. Shen, SOLOv2: dynamic and fast instance segmentation, in: H. Larochelle, M. Ranzato, R. Hadsell, M.F. Balcan, H. Lin (Eds.), Advances in Neural Information Processing Systems (NeurIPS), 2020, pp. 17721-17732.
[6] D. Bolya, C. Zhou, F. Xiao, Y.J. Lee, YOLACT: real-time instance segmentation, in: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2019, pp. 9158-9166.
[7] E. Mohamed, A. Shaker, A. El-Sallab, M. Hadhoud, INSTA-YOLO: real-time instance segmentation, arXiv:2102.0677, 2021.
[8] G. Jocher, YOLO by Ultralytics (version 5.7.0), GitHub, 2022. https://github.com/ ultralytics/yolov5.
[9] G. Jocher, A. Chaurasia, J. Qiu, YOLO by Ultralytics (version 8.0.0), GitHub, 2023. https://github.com/ultralytics/ultralytics.
[10] C.-Y. Wang, H.-Y.M. Liao, Y.-H. Wu, P.-Y. Chen, J.-W. Hsieh, I.-H. Yeh, CSPNet: A new backbone that can enhance learning capability of CNN, in: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), 2020, pp. 1571-1580.
[11] Ultralytics, Ultralytics YOLOv5 Architecture, Ultralytics, 2023. https://docs.ultr alytics.com/yolov5/tutorials/architecture_description.
[12] A. Bochkovskiy, C.Y. Wang, H.Y.M. Liao, YOLOv4: Optimal speed and accuracy of object detection, arXiv:2004.10934, 2020.
[13] T.-Y. Lin, P. Dollár, R. Girshick, K. He, B. Hariharan, S. Belongie, Feature pyramid networks for object detection, in: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2017, pp. 2117-2125.
[14] Y.-F. Zhang, W. Ren, Z. Zhang, Z. Jia, L. Wang, T. Tan, Focal and efficient IOU loss for accurate bounding box regression, Neurocomputing 506 (2022) 146-157.
[15] N. Bodla, B. Singh, R. Chellappa, L.S. Davis, Soft-NMS-improving object detection with one line of code, in: Proceedings of the IEEE International Conference on Computer Vision (ICCV), 2017, pp. 5562-5570.
[16] S. Liu, L. Qi, H. Qin, J. Shi, J. Jia, Path aggregation network for instance segmentation, in: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2018, pp. 8759-8768.
[17] R. Hollandi, N. Moshkov, L. Paavolainen, E. Tasnadi, F. Piccinini, P. Horvath, Nucleus segmentation: towards automated solutions, Trends Cell Biol. 32 (4) (2021) 295-310.
[18] J. Johnson, Adapting mask-RCNN for automatic nucleus segmentation, arXiv: 1805.00500, 2018.
[19] H. Jung, B. Lodhi, J. Kang, An automatic nuclei segmentation method based on deep convolutional neural networks for histopathology images, BMC Biomed. Eng. 1 (2019) 24.
[20] S. Fujita, X.-H. Han, Cell detection and segmentation in microscopy images with improved mask R-CNN, in: I. Sato, B. Han (Eds.), Computer Vision – ACCV 2020 Workshops, 2020, pp. 58-70.
[21] B. Bancher, A. Mahbod, I. Ellinger, Improving mask R-CNN for nuclei instance segmentation in hematoxylin & eosin-stained histological images, in: M. Atzori, N. Burlutskiy, F. Ciompi, Z. Li, F. Minhas, H. Müller, et al. (Eds.), Proceedings of the MICCAI Workshop on Computational Pathology, PMLR 156, 2024, pp. 20-35.
[22] J. Yi, P. Wu, M. Jiang, Q. Huang, D.J. Hoeppner, D.N. Metaxas, Attentive neural cell instance segmentation, Med. Image Anal. 55 (2019) 228-240.
[23] Z. Cheng, A. Qu, A fast and accurate algorithm for nuclei instance segmentation in microscopy images, IEEE Access 8 (2020) 158679-158689.
[24] W. Liu, D. Anguelov, D. Erhan, C. Szegedy, S. Reed, C. Fu, et al., SSD: Single shot multibox detector, in: B. Leibe, J. Matas, N. Sebe, M. Welling (Eds.), Computer Vision – ECCV 2016, Part I, 2016, pp. 21-37.
[25] A. Mahbod, G. Schaefer, G. Dorffner, S. Hatamikia, R. Ecker, I. Ellinger, A dual decoder U-net-based model for nuclei instance segmentation in hematoxylin and eosin-stained histological images, Front. Med. 9 (2022) 978146.
[26] O. Ronneberger, P. Fischer, T. Brox, U-Net: Convolutional networks for biomedical image segmentation, in: N. Navab, J. Hornegger, W.M. Wells, A.F. Frangi (Eds.), Medical Image Computing and Computer-Assisted Intervention – MICCAI, Part III, 2015, pp. 234-241.
[27] T. Konopczyński, R. Heiman, P. Woźnicki, P. Gniewek, M.-C. Duvernoy, O. Hallatschek, et al., Instance segmentation of densely packed cells using a hybrid model of U-Net and mask R-CNN, in: L. Rutkowski, R. Scherer, M. Korytkowski, W. Pedrycz, R. Tadeusiewicz, J.M. Zurada (Eds.), Artificial Intelligence and Soft Computing (ICAISC), Part I, 2020, pp. 626-635.
[28] J. Wang, Z. Zhang, M. Wu, Y. Ye, S. Wang, Y. Cao, et al., Improved BlendMask: nuclei instance segmentation for medical microscopy images, IET Image Process. 17 (7) (2023) 2284-2296.
[29] J. Hu, L. Shen, G. Sun, Squeeze-and-excitation networks, in: Proceedings of IEEE/ CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2018, pp. 7132-7141.
[30] Z. Shang, X. Wang, Y. Jiang, Z. Li, J. Ning, Identifying rumen protozoa in microscopic images of ruminant with improved YOLACT instance segmentation, Biosyst. Eng. 215 (2022) 156-169.
[31] Y. Wang, Z. Ouyang, R. Han, Z. Yin, Z. Yang, YOLOMask: Real-time instance segmentation with integrating YOLOv5 and OrienMask, in: Proceedings of the IEEE 22nd International Conference on Communication Technology (ICCT), 2022, pp. 1646-1650.
[32] W. Yang, S. Chen, G. Chen, Q. Shi, PR-YOLO: Improved YOLO for fast protozoa classification and segmentation, Res. Square Preprint (2024), https://doi.org/ 10.21203/rs.3.rs-3199595/v1.
[33] X. Cao, Y. Su, X. Geng, Y. Wang, YOLO-SF: YOLO for fire segmentation detection, IEEE Access 11 (2023) 111079-111092.
[34] C.-Y. Wang, A. Bochkovskiy, H.-Y.M. Liao, YOLOv7: Trainable bag-of-freebies sets new state-of-the-art for real-time object detectors, in: Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2023, pp. 7464-7475.
[35] S. Woo, J. Park, J.-Y. Lee, I.S. Kweon, CBAM: Convolutional block attention module, in: V. Ferrari, M. Hebert, C. Sminchisescu, Y. Weiss (Eds.), Computer Vision – ECCV 2018, Part VII, 2018, pp. 3-19.
[36] O.M. Lawal, YOLOv5-LiNet: a lightweight network for fruits instance segmentation, PLoS One 18 (3) (2023) e0282297.
[37] M. Yasir, L. Zhan, S. Liu, J. Wan, M.S. Hossain, A.T. Isiacik Çolak, et al., Instance segmentation ship detection based on improved Yolov7 using complex background SAR images, Front. Mar. Sci. 10 (2023) 1113669.
[38] H. Liu, W. Xiong, Y. Zhang, YOLO-CORE: contour regression for efficient instance segmentation, Mach. Intell. Res. 20 (2023) 716-728.
[39] J. Hua, T. Hao, L. Zeng, G. Yu, YOLOMask, an instance segmentation algorithm based on complementary fusion network, Math 9 (15) (2021) 1766.
[40] B. Bai, J. Tian, T. Wang, S. Luo, S. Lyu, YUSEG: Yolo and Unet is all you need for cell instance segmentation, in: NeurIPS 2022 Weakly Supervised Cell Segmentation in Multi-modality High-Resolution Microscopy Images, 2022.
[41] T. Lindeberg, Scale-Space Theory in Computer Vision, Springer, Cham, 1994, pp. 10-11.
[42] D.J. Lowe, Distinctive image features from scale-invariant keypoints, Int. J. Comput. Vis. 60 (2004) 91-110.
[43] D. Tran, L. Bourdev, R. Fergus, L. Torresani, M. Paluri, Learning spatiotemporal features with 3D convolutional networks, in: Proceedings of IEEE International Conference on Computer Vision (ICCV), 2015, pp. 4489-4497.
[44] H.-J. Park, J.-W. Kang, B.-G. Kim, ssFPN: scale sequence (
[45] O. Rukundo, H. Cao, Nearest neighbor value interpolation, Int. J. Adv. Comput. Sci. Appl. 3 (4) (2012) 25-30.
[46] S. Elfwing, E. Uchibe, K. Doya, Sigmoid-weighted linear units for neural network function approximation in reinforcement learning, Neural Netw. 107 (2018) 3-11.
[47] H. Rezatofighi, N. Tsoi, J. Gwak, A. Sadeghian, I. Reid, S. Savarese, Generalized intersection over union: a metric and a loss for bounding box pegression, in: Proceedings of IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2019, pp. 658-666.
[48] Z. Zheng, P. Wang, W. Liu, J. Li, R. Ye, D. Ren, Distance-IoU loss: faster and better learning for bounding box regression, Proc. AAAI Conf. Artific. Intellig. 34 (07) (2020) 12993-13000.
[49] A. Neubeck, L. Van Gool, Efficient non-maximum suppression, in: Proceedings of the 18th International Conference on Pattern Recognition (ICPR), 2006, pp. 850-855.
[50] A. Goodman, A. Carpenter, E. Park, J. Lefman, J. BoozAllen, K. Thomas, et al., 2018 Data Science Bowl, Kaggle, 2018. https://kaggle.com/competitions/data-s cience-bowl-2018.
[51] CBI, Breast Cancer Cell, UCSB, 2008. https://bioimage.ucsb.edu/research/bio-seg mentation.
[52] OpenMMLab, Mask-rcnn_swin-t, GitHub, 2022. https://github.com/open-mm lab/mmdetection/tree/main/configs/swin.
[53] Q. Hou, D. Zhou, J. Feng, Coordinate attention for efficient mobile network design, in: Proceedings of IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), 2021, pp. 13708-13717.
[54] P. Wu, H. Li, N. Zeng, F. Li, FMD-Yolo: an efficient face mask detection method for COVID-19 prevention and control in public, Image Vis. Comput. 117 (2022) 104341.
[55] X. Guo, Z. Wang, P. Wu, Y. Li, F.E. Alsaadi, N. Zeng, ELTS-net: an enhanced liver tumor segmentation network with augmented receptive field and global contextual information, Comput. Biol. Med. 169 (2024) 107879.
[56] L. Wang, Y. Jiao, Y. Diao, N. Zeng, R. Yu, A novel approach combined transfer learning and deep learning to predict TMB from histology image, Pattern Recogn. Lett. 135 (2020) 244-248.
- *Corresponding author.
E-mail address:ting.cheeming@monash.edu(C.-M.Ting).
